Testing hierarchical pathway kinetics with residue data on cyantraniliprole
Johannes Ranke
Last change on 13 February 2023, last compiled on 12 Mai 2025
Source:vignettes/prebuilt/2022_cyan_pathway.rmd
2022_cyan_pathway.rmdIntroduction
The purpose of this document is to test demonstrate how nonlinear hierarchical models (NLHM) based on the parent degradation models SFO, FOMC, DFOP and HS, with serial formation of two or more metabolites can be fitted with the mkin package.
It was assembled in the course of work package 1.2 of Project Number 173340 (Application of nonlinear hierarchical models to the kinetic evaluation of chemical degradation data) of the German Environment Agency carried out in 2022 and 2023.
The mkin package is used in version 1.2.10 which is currently under
development. The newly introduced functionality that is used here is a
simplification of excluding random effects for a set of fits based on a
related set of fits with a reduced model, and the documentation of the
starting parameters of the fit, so that all starting parameters of
saem fits are now listed in the summary. The
saemix package is used as a backend for fitting the NLHM,
but is also loaded to make the convergence plot function available.
This document is processed with the knitr package, which
also provides the kable function that is used to improve
the display of tabular data in R markdown documents. For parallel
processing, the parallel package is used.
library(mkin)
library(knitr)
library(saemix)
library(parallel)
n_cores <- detectCores()
# We need to start a new cluster after defining a compiled model that is
# saved as a DLL to the user directory, therefore we define a function
# This is used again after defining the pathway model
start_cluster <- function(n_cores) {
if (Sys.info()["sysname"] == "Windows") {
ret <- makePSOCKcluster(n_cores)
} else {
ret <- makeForkCluster(n_cores)
}
return(ret)
}
cl <- start_cluster(n_cores)Test data
The example data are taken from the final addendum to the DAR from
2014 and are distributed with the mkin package. Residue data and time
step normalisation factors are read in using the function
read_spreadsheet from the mkin package. This function also
performs the time step normalisation.
data_file <- system.file(
"testdata", "cyantraniliprole_soil_efsa_2014.xlsx",
package = "mkin")
cyan_ds <- read_spreadsheet(data_file, parent_only = FALSE)The following tables show the covariate data and the 5 datasets that were read in from the spreadsheet file.
| pH | |
|---|---|
| Nambsheim | 7.90 |
| Tama | 6.20 |
| Gross-Umstadt | 7.04 |
| Sassafras | 4.62 |
| Lleida | 8.05 |
for (ds_name in names(cyan_ds)) {
print(
kable(mkin_long_to_wide(cyan_ds[[ds_name]]),
caption = paste("Dataset", ds_name),
booktabs = TRUE, row.names = FALSE))
cat("\n\\clearpage\n")
}| time | cyan | JCZ38 | J9C38 | JSE76 | J9Z38 |
|---|---|---|---|---|---|
| 0.000000 | 105.79 | NA | NA | NA | NA |
| 3.210424 | 77.26 | 7.92 | 11.94 | 5.58 | 9.12 |
| 7.490988 | 57.13 | 15.46 | 16.58 | 12.59 | 11.74 |
| 17.122259 | 37.74 | 15.98 | 13.36 | 26.05 | 10.77 |
| 23.543105 | 31.47 | 6.05 | 14.49 | 34.71 | 4.96 |
| 43.875788 | 16.74 | 6.07 | 7.57 | 40.38 | 6.52 |
| 67.418893 | 8.85 | 10.34 | 6.39 | 30.71 | 8.90 |
| 107.014116 | 5.19 | 9.61 | 1.95 | 20.41 | 12.93 |
| 129.487080 | 3.45 | 6.18 | 1.36 | 21.78 | 6.99 |
| 195.835832 | 2.15 | 9.13 | 0.95 | 16.29 | 7.69 |
| 254.693596 | 1.92 | 6.92 | 0.20 | 13.57 | 7.16 |
| 321.042348 | 2.26 | 7.02 | NA | 11.12 | 8.66 |
| 383.110535 | NA | 5.05 | NA | 10.64 | 5.56 |
| 0.000000 | 105.57 | NA | NA | NA | NA |
| 3.210424 | 78.88 | 12.77 | 11.94 | 5.47 | 9.12 |
| 7.490988 | 59.94 | 15.27 | 16.58 | 13.60 | 11.74 |
| 17.122259 | 39.67 | 14.26 | 13.36 | 29.44 | 10.77 |
| 23.543105 | 30.21 | 16.07 | 14.49 | 35.90 | 4.96 |
| 43.875788 | 18.06 | 9.44 | 7.57 | 42.30 | 6.52 |
| 67.418893 | 8.54 | 5.78 | 6.39 | 34.70 | 8.90 |
| 107.014116 | 7.26 | 4.54 | 1.95 | 23.33 | 12.93 |
| 129.487080 | 3.60 | 4.22 | 1.36 | 23.56 | 6.99 |
| 195.835832 | 2.84 | 3.05 | 0.95 | 16.21 | 7.69 |
| 254.693596 | 2.00 | 2.90 | 0.20 | 15.53 | 7.16 |
| 321.042348 | 1.79 | 0.94 | NA | 9.80 | 8.66 |
| 383.110535 | NA | 1.82 | NA | 9.49 | 5.56 |
| time | cyan | JCZ38 | J9Z38 | JSE76 |
|---|---|---|---|---|
| 0.000000 | 106.14 | NA | NA | NA |
| 2.400833 | 93.47 | 6.46 | 2.85 | NA |
| 5.601943 | 88.39 | 10.86 | 4.65 | 3.85 |
| 12.804442 | 72.29 | 11.97 | 4.91 | 11.24 |
| 17.606108 | 65.79 | 13.11 | 6.63 | 13.79 |
| 32.811382 | 53.16 | 11.24 | 8.90 | 23.40 |
| 50.417490 | 44.01 | 11.34 | 9.98 | 29.56 |
| 80.027761 | 33.23 | 8.82 | 11.31 | 35.63 |
| 96.833591 | 40.68 | 5.94 | 8.32 | 29.09 |
| 146.450803 | 20.65 | 4.49 | 8.72 | 36.88 |
| 190.466072 | 17.71 | 4.66 | 11.10 | 40.97 |
| 240.083284 | 14.86 | 2.27 | 11.62 | 40.11 |
| 286.499386 | 12.02 | NA | 10.73 | 42.58 |
| 0.000000 | 109.11 | NA | NA | NA |
| 2.400833 | 96.84 | 5.52 | 2.04 | 2.02 |
| 5.601943 | 85.29 | 9.65 | 2.99 | 4.39 |
| 12.804442 | 73.68 | 12.48 | 5.05 | 11.47 |
| 17.606108 | 64.89 | 12.44 | 6.29 | 15.00 |
| 32.811382 | 52.27 | 10.86 | 7.65 | 23.30 |
| 50.417490 | 42.61 | 10.54 | 9.37 | 31.06 |
| 80.027761 | 34.29 | 10.02 | 9.04 | 37.87 |
| 96.833591 | 30.50 | 6.34 | 8.14 | 33.97 |
| 146.450803 | 19.21 | 6.29 | 8.52 | 26.15 |
| 190.466072 | 17.55 | 5.81 | 9.89 | 32.08 |
| 240.083284 | 13.22 | 5.99 | 10.79 | 40.66 |
| 286.499386 | 11.09 | 6.05 | 8.82 | 42.90 |
| time | cyan | JCZ38 | J9Z38 | JSE76 |
|---|---|---|---|---|
| 0.0000000 | 103.03 | NA | NA | NA |
| 2.1014681 | 87.85 | 4.79 | 3.26 | 0.62 |
| 4.9034255 | 77.35 | 8.05 | 9.89 | 1.32 |
| 10.5073404 | 69.33 | 9.74 | 12.32 | 4.74 |
| 21.0146807 | 55.65 | 14.57 | 13.59 | 9.84 |
| 31.5220211 | 49.03 | 14.66 | 16.71 | 12.32 |
| 42.0293615 | 41.86 | 15.97 | 13.64 | 15.53 |
| 63.0440422 | 34.88 | 18.20 | 14.12 | 22.02 |
| 84.0587230 | 28.26 | 15.64 | 14.06 | 25.60 |
| 0.0000000 | 104.05 | NA | NA | NA |
| 2.1014681 | 85.25 | 2.68 | 7.32 | 0.69 |
| 4.9034255 | 77.22 | 7.28 | 8.37 | 1.45 |
| 10.5073404 | 65.23 | 10.73 | 10.93 | 4.74 |
| 21.0146807 | 57.78 | 12.29 | 14.80 | 9.05 |
| 31.5220211 | 54.83 | 14.05 | 12.01 | 11.05 |
| 42.0293615 | 45.17 | 12.12 | 17.89 | 15.71 |
| 63.0440422 | 34.83 | 12.90 | 15.86 | 22.52 |
| 84.0587230 | 26.59 | 14.28 | 14.91 | 28.48 |
| 0.0000000 | 104.62 | NA | NA | NA |
| 0.8145225 | 97.21 | NA | 4.00 | NA |
| 1.9005525 | 89.64 | 3.59 | 5.24 | NA |
| 4.0726125 | 87.90 | 4.10 | 9.58 | NA |
| 8.1452251 | 86.90 | 5.96 | 9.45 | NA |
| 12.2178376 | 74.74 | 7.83 | 15.03 | 5.33 |
| 16.2904502 | 74.13 | 8.84 | 14.41 | 5.10 |
| 24.4356753 | 65.26 | 11.84 | 18.33 | 6.71 |
| 32.5809004 | 57.70 | 12.74 | 19.93 | 9.74 |
| 0.0000000 | 101.94 | NA | NA | NA |
| 0.8145225 | 99.94 | NA | NA | NA |
| 1.9005525 | 94.87 | NA | 4.56 | NA |
| 4.0726125 | 86.96 | 6.75 | 6.90 | NA |
| 8.1452251 | 80.51 | 10.68 | 7.43 | 2.58 |
| 12.2178376 | 78.38 | 10.35 | 9.46 | 3.69 |
| 16.2904502 | 70.05 | 13.73 | 9.27 | 7.18 |
| 24.4356753 | 61.28 | 12.57 | 13.28 | 13.19 |
| 32.5809004 | 52.85 | 12.67 | 12.95 | 13.69 |
| time | cyan | JCZ38 | J9Z38 | JSE76 |
|---|---|---|---|---|
| 0.000000 | 102.17 | NA | NA | NA |
| 2.216719 | 95.49 | 1.11 | 0.10 | 0.83 |
| 5.172343 | 83.35 | 6.43 | 2.89 | 3.30 |
| 11.083593 | 78.18 | 10.00 | 5.59 | 0.81 |
| 22.167186 | 70.44 | 17.21 | 4.23 | 1.09 |
| 33.250779 | 68.00 | 20.45 | 5.86 | 1.17 |
| 44.334371 | 59.64 | 24.64 | 3.17 | 2.72 |
| 66.501557 | 50.73 | 27.50 | 6.19 | 1.27 |
| 88.668742 | 45.65 | 32.77 | 5.69 | 4.54 |
| 0.000000 | 100.43 | NA | NA | NA |
| 2.216719 | 95.34 | 3.21 | 0.14 | 0.46 |
| 5.172343 | 84.38 | 5.73 | 4.75 | 0.62 |
| 11.083593 | 78.50 | 11.89 | 3.99 | 0.73 |
| 22.167186 | 71.17 | 17.28 | 4.39 | 0.66 |
| 33.250779 | 59.41 | 18.73 | 11.85 | 2.65 |
| 44.334371 | 64.57 | 22.93 | 5.13 | 2.01 |
| 66.501557 | 49.08 | 33.39 | 5.67 | 3.63 |
| 88.668742 | 40.41 | 39.60 | 5.93 | 6.17 |
| time | cyan | JCZ38 | J9Z38 | JSE76 |
|---|---|---|---|---|
| 0.000000 | 102.71 | NA | NA | NA |
| 2.821051 | 79.11 | 5.70 | 8.07 | 0.97 |
| 6.582451 | 70.03 | 7.17 | 11.31 | 4.72 |
| 14.105253 | 50.93 | 10.25 | 14.84 | 9.95 |
| 28.210505 | 33.43 | 10.40 | 14.82 | 24.06 |
| 42.315758 | 24.69 | 9.75 | 16.38 | 29.38 |
| 56.421010 | 22.99 | 10.06 | 15.51 | 29.25 |
| 84.631516 | 14.63 | 5.63 | 14.74 | 31.04 |
| 112.842021 | 12.43 | 4.17 | 13.53 | 33.28 |
| 0.000000 | 99.31 | NA | NA | NA |
| 2.821051 | 82.07 | 6.55 | 5.60 | 1.12 |
| 6.582451 | 70.65 | 7.61 | 8.01 | 3.21 |
| 14.105253 | 53.52 | 11.48 | 10.82 | 12.24 |
| 28.210505 | 35.60 | 11.19 | 15.43 | 23.53 |
| 42.315758 | 34.26 | 11.09 | 13.26 | 27.42 |
| 56.421010 | 21.79 | 4.80 | 18.30 | 30.20 |
| 84.631516 | 14.06 | 6.30 | 16.35 | 32.32 |
| 112.842021 | 11.51 | 5.57 | 12.64 | 32.51 |
Parent only evaluations
As the pathway fits have very long run times, evaluations of the parent data are performed first, in order to determine for each hierarchical parent degradation model which random effects on the degradation model parameters are ill-defined.
cyan_sep_const <- mmkin(c("SFO", "FOMC", "DFOP", "SFORB", "HS"),
cyan_ds, quiet = TRUE, cores = n_cores)
cyan_sep_tc <- update(cyan_sep_const, error_model = "tc")
cyan_saem_full <- mhmkin(list(cyan_sep_const, cyan_sep_tc))
status(cyan_saem_full) |> kable()| const | tc | |
|---|---|---|
| SFO | OK | OK |
| FOMC | OK | OK |
| DFOP | OK | OK |
| SFORB | OK | OK |
| HS | OK | OK |
All fits converged successfully.
| const | tc | |
|---|---|---|
| SFO | sd(cyan_0) | sd(cyan_0) |
| FOMC | sd(log_beta) | sd(cyan_0) |
| DFOP | sd(cyan_0) | sd(cyan_0), sd(log_k1) |
| SFORB | sd(cyan_free_0) | sd(cyan_free_0), sd(log_k_cyan_free_bound) |
| HS | sd(cyan_0) | sd(cyan_0) |
In almost all models, the random effect for the initial concentration of the parent compound is ill-defined. For the biexponential models DFOP and SFORB, the random effect of one additional parameter is ill-defined when the two-component error model is used.
| npar | AIC | BIC | Lik | |
|---|---|---|---|---|
| SFO const | 5 | 833.9 | 832.0 | -412.0 |
| SFO tc | 6 | 831.6 | 829.3 | -409.8 |
| FOMC const | 7 | 709.1 | 706.4 | -347.6 |
| FOMC tc | 8 | 689.2 | 686.1 | -336.6 |
| DFOP const | 9 | 703.0 | 699.5 | -342.5 |
| SFORB const | 9 | 701.3 | 697.8 | -341.7 |
| HS const | 9 | 718.6 | 715.1 | -350.3 |
| DFOP tc | 10 | 703.1 | 699.2 | -341.6 |
| SFORB tc | 10 | 700.0 | 696.1 | -340.0 |
| HS tc | 10 | 716.7 | 712.8 | -348.3 |
Model comparison based on AIC and BIC indicates that the two-component error model is preferable for all parent models with the exception of DFOP. The lowest AIC and BIC values are are obtained with the FOMC model, followed by SFORB and DFOP.
stopCluster(cl)Pathway fits
Evaluations with pathway established previously
To test the technical feasibility of coupling the relevant parent
degradation models with different transformation pathway models, a list
of mkinmod models is set up below. As in the EU evaluation,
parallel formation of metabolites JCZ38 and J9Z38 and secondary
formation of metabolite JSE76 from JCZ38 is used.
if (!dir.exists("cyan_dlls")) dir.create("cyan_dlls")
cyan_path_1 <- list(
sfo_path_1 = mkinmod(
cyan = mkinsub("SFO", c("JCZ38", "J9Z38")),
JCZ38 = mkinsub("SFO", "JSE76"),
J9Z38 = mkinsub("SFO"),
JSE76 = mkinsub("SFO"), quiet = TRUE,
name = "sfo_path_1", dll_dir = "cyan_dlls", overwrite = TRUE),
fomc_path_1 = mkinmod(
cyan = mkinsub("FOMC", c("JCZ38", "J9Z38")),
JCZ38 = mkinsub("SFO", "JSE76"),
J9Z38 = mkinsub("SFO"),
JSE76 = mkinsub("SFO"), quiet = TRUE,
name = "fomc_path_1", dll_dir = "cyan_dlls", overwrite = TRUE),
dfop_path_1 = mkinmod(
cyan = mkinsub("DFOP", c("JCZ38", "J9Z38")),
JCZ38 = mkinsub("SFO", "JSE76"),
J9Z38 = mkinsub("SFO"),
JSE76 = mkinsub("SFO"), quiet = TRUE,
name = "dfop_path_1", dll_dir = "cyan_dlls", overwrite = TRUE),
sforb_path_1 = mkinmod(
cyan = mkinsub("SFORB", c("JCZ38", "J9Z38")),
JCZ38 = mkinsub("SFO", "JSE76"),
J9Z38 = mkinsub("SFO"),
JSE76 = mkinsub("SFO"), quiet = TRUE,
name = "sforb_path_1", dll_dir = "cyan_dlls", overwrite = TRUE),
hs_path_1 = mkinmod(
cyan = mkinsub("HS", c("JCZ38", "J9Z38")),
JCZ38 = mkinsub("SFO", "JSE76"),
J9Z38 = mkinsub("SFO"),
JSE76 = mkinsub("SFO"), quiet = TRUE,
name = "hs_path_1", dll_dir = "cyan_dlls", overwrite = TRUE)
)
cl_path_1 <- start_cluster(n_cores)To obtain suitable starting values for the NLHM fits, separate pathway fits are performed for all datasets.
f_sep_1_const <- mmkin(
cyan_path_1,
cyan_ds,
error_model = "const",
cluster = cl_path_1,
quiet = TRUE)
status(f_sep_1_const) |> kable()| Nambsheim | Tama | Gross-Umstadt | Sassafras | Lleida | |
|---|---|---|---|---|---|
| sfo_path_1 | OK | OK | OK | C | OK |
| fomc_path_1 | OK | OK | OK | OK | OK |
| dfop_path_1 | OK | OK | OK | OK | OK |
| sforb_path_1 | OK | OK | OK | OK | OK |
| hs_path_1 | C | C | C | C | C |
| Nambsheim | Tama | Gross-Umstadt | Sassafras | Lleida | |
|---|---|---|---|---|---|
| sfo_path_1 | OK | OK | OK | OK | OK |
| fomc_path_1 | OK | OK | OK | OK | OK |
| dfop_path_1 | OK | OK | OK | OK | OK |
| sforb_path_1 | OK | OK | OK | OK | OK |
| hs_path_1 | C | OK | C | OK | C |
Most separate fits converged successfully. The biggest convergence problems are seen when using the HS model with constant variance.
For the hierarchical pathway fits, those random effects that could not be quantified in the corresponding parent data analyses are excluded.
In the code below, the output of the illparms function
for the parent only fits is used as an argument
no_random_effect to the mhmkin function. The
possibility to do so was introduced in mkin version 1.2.2
which is currently under development.
f_saem_1 <- mhmkin(list(f_sep_1_const, f_sep_1_tc),
no_random_effect = illparms(cyan_saem_full),
cluster = cl_path_1)| const | tc | |
|---|---|---|
| sfo_path_1 | FO | Fth, FO |
| fomc_path_1 | OK | Fth, FO |
| dfop_path_1 | Fth, FO | Fth, FO |
| sforb_path_1 | Fth, FO | Fth, FO |
| hs_path_1 | FO | E |
The status information from the individual fits shows that all fits
completed successfully. The matrix entries Fth and FO indicate that the
Fisher Information Matrix could not be inverted for the fixed effects
(theta) and the random effects (Omega), respectively. For the affected
fits, ill-defined parameters cannot be determined using the
illparms function, because it relies on the Fisher
Information Matrix.
| const | tc | |
|---|---|---|
| sfo_path_1 | NA | NA |
| fomc_path_1 | sd(log_k_J9Z38), sd(f_cyan_ilr_2), sd(f_JCZ38_qlogis) | NA |
| dfop_path_1 | NA | NA |
| sforb_path_1 | NA | NA |
| hs_path_1 | NA | E |
The model comparisons below suggest that the pathway fits using DFOP or SFORB for the parent compound provide the best fit.
| npar | AIC | BIC | Lik | |
|---|---|---|---|---|
| sfo_path_1 const | 16 | 2693.0 | 2686.8 | -1330.5 |
| fomc_path_1 const | 18 | 2427.9 | 2420.9 | -1196.0 |
| dfop_path_1 const | 20 | 2403.2 | 2395.4 | -1181.6 |
| sforb_path_1 const | 20 | 2401.4 | 2393.6 | -1180.7 |
| hs_path_1 const | 20 | 2427.2 | 2419.4 | -1193.6 |
| npar | AIC | BIC | Lik | |
|---|---|---|---|---|
| sfo_path_1 const | 16 | 2693.0 | 2686.8 | -1330.5 |
| sfo_path_1 tc | 17 | 2657.6 | 2651.0 | -1311.8 |
| fomc_path_1 const | 18 | 2427.9 | 2420.9 | -1196.0 |
| fomc_path_1 tc | 19 | 2423.6 | 2416.2 | -1192.8 |
| dfop_path_1 const | 20 | 2403.2 | 2395.4 | -1181.6 |
| sforb_path_1 const | 20 | 2401.4 | 2393.6 | -1180.7 |
| dfop_path_1 tc | 20 | 2398.0 | 2390.1 | -1179.0 |
| sforb_path_1 tc | 20 | 2399.9 | 2392.1 | -1180.0 |
For these two parent model, successful fits are shown below. Plots of the fits with the other parent models are shown in the Appendix.
plot(f_saem_1[["dfop_path_1", "tc"]])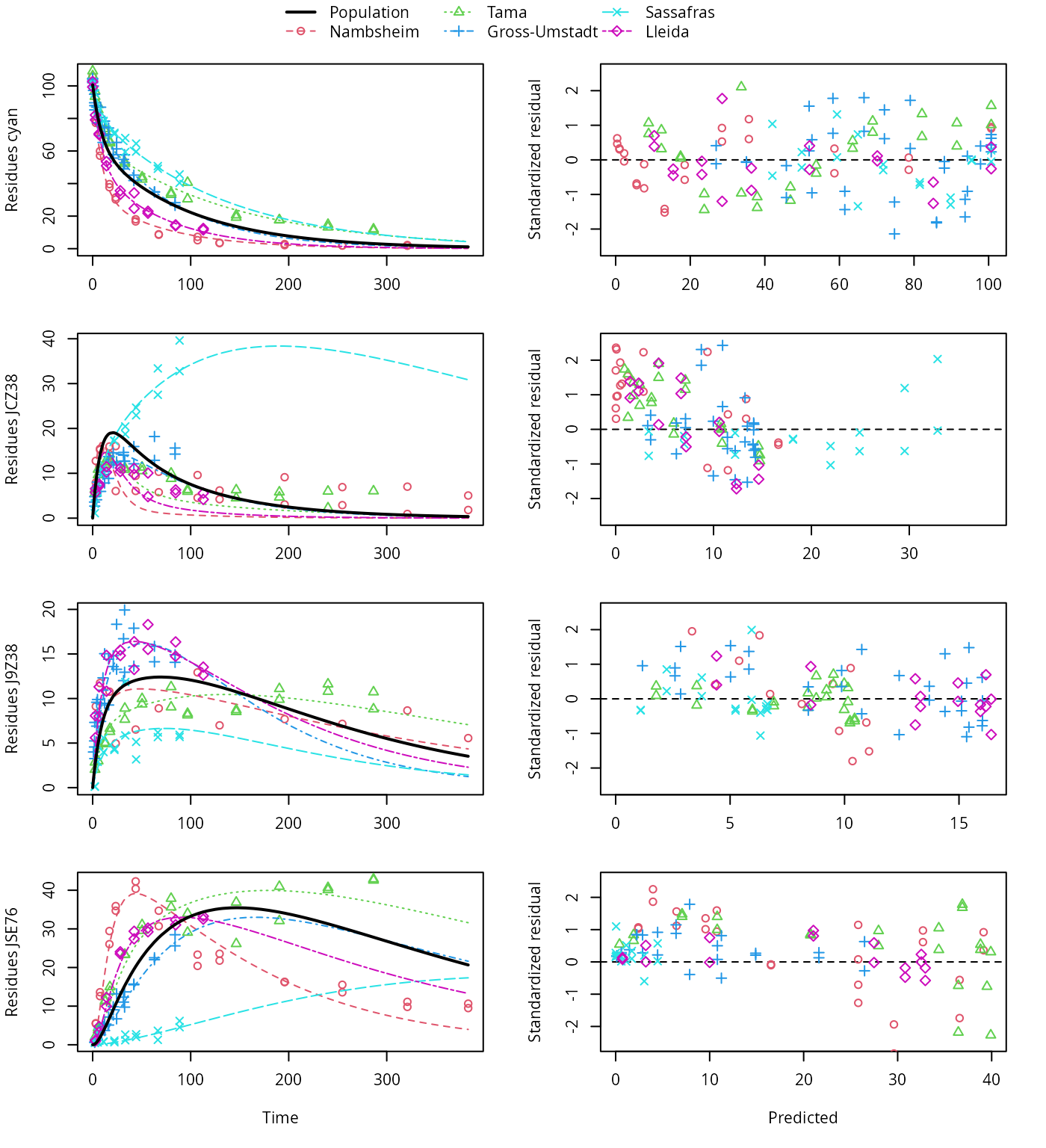
DFOP pathway fit with two-component error
plot(f_saem_1[["sforb_path_1", "tc"]])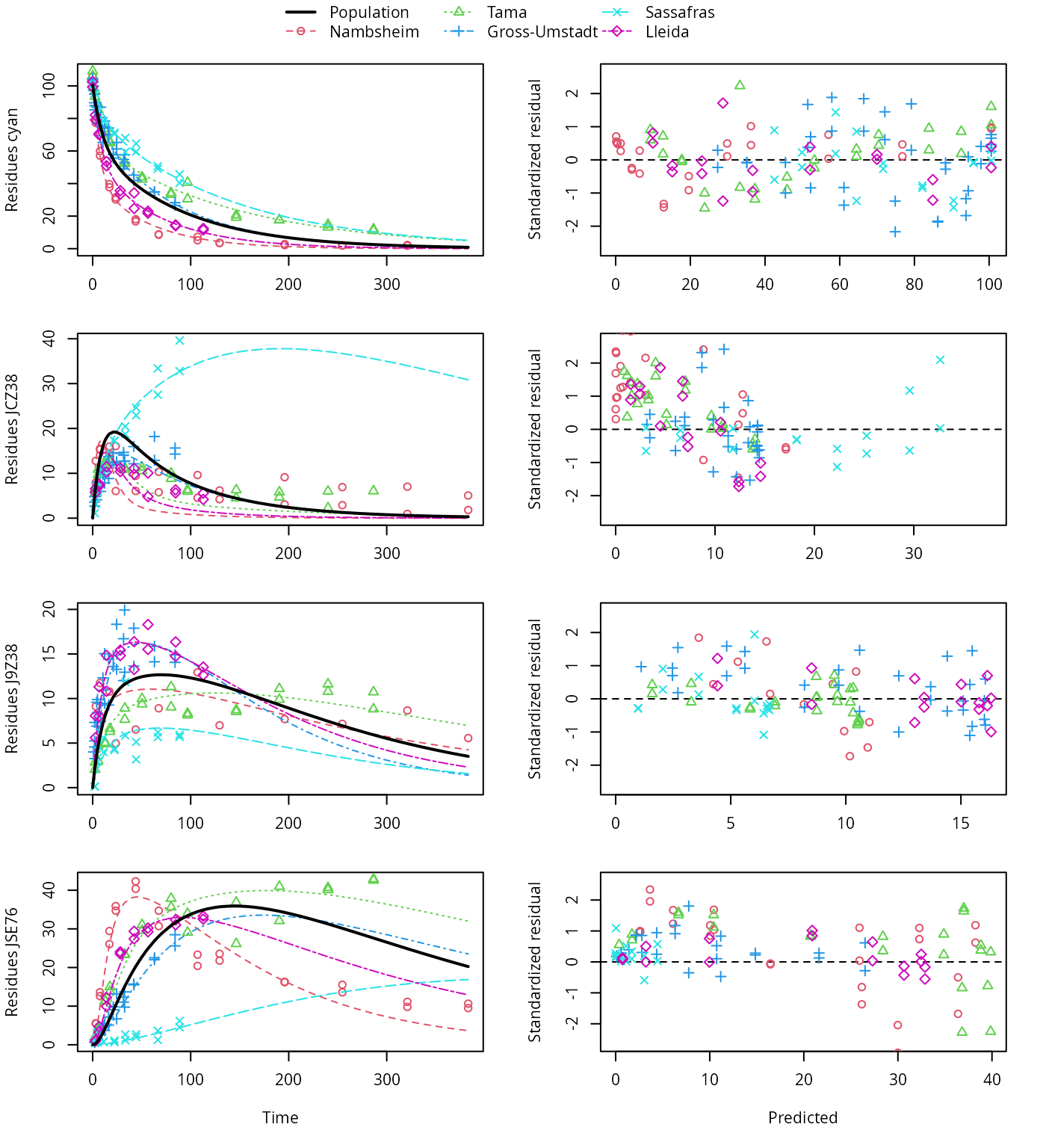
SFORB pathway fit with two-component error
A closer graphical analysis of these Figures shows that the residues of transformation product JCZ38 in the soils Tama and Nambsheim observed at later time points are strongly and systematically underestimated.
stopCluster(cl_path_1)Alternative pathway fits
To improve the fit for JCZ38, a back-reaction from JSE76 to JCZ38 was introduced in an alternative version of the transformation pathway, in analogy to the back-reaction from K5A78 to K5A77. Both pairs of transformation products are pairs of an organic acid with its corresponding amide (Addendum 2014, p. 109). As FOMC provided the best fit for the parent, and the biexponential models DFOP and SFORB provided the best initial pathway fits, these three parent models are used in the alternative pathway fits.
cyan_path_2 <- list(
fomc_path_2 = mkinmod(
cyan = mkinsub("FOMC", c("JCZ38", "J9Z38")),
JCZ38 = mkinsub("SFO", "JSE76"),
J9Z38 = mkinsub("SFO"),
JSE76 = mkinsub("SFO", "JCZ38"),
name = "fomc_path_2", quiet = TRUE,
dll_dir = "cyan_dlls",
overwrite = TRUE
),
dfop_path_2 = mkinmod(
cyan = mkinsub("DFOP", c("JCZ38", "J9Z38")),
JCZ38 = mkinsub("SFO", "JSE76"),
J9Z38 = mkinsub("SFO"),
JSE76 = mkinsub("SFO", "JCZ38"),
name = "dfop_path_2", quiet = TRUE,
dll_dir = "cyan_dlls",
overwrite = TRUE
),
sforb_path_2 = mkinmod(
cyan = mkinsub("SFORB", c("JCZ38", "J9Z38")),
JCZ38 = mkinsub("SFO", "JSE76"),
J9Z38 = mkinsub("SFO"),
JSE76 = mkinsub("SFO", "JCZ38"),
name = "sforb_path_2", quiet = TRUE,
dll_dir = "cyan_dlls",
overwrite = TRUE
)
)
cl_path_2 <- start_cluster(n_cores)
f_sep_2_const <- mmkin(
cyan_path_2,
cyan_ds,
error_model = "const",
cluster = cl_path_2,
quiet = TRUE)
status(f_sep_2_const) |> kable()| Nambsheim | Tama | Gross-Umstadt | Sassafras | Lleida | |
|---|---|---|---|---|---|
| fomc_path_2 | OK | OK | OK | C | OK |
| dfop_path_2 | OK | OK | OK | C | OK |
| sforb_path_2 | OK | OK | OK | OK | OK |
Using constant variance, separate fits converge with the exception of the fits to the Sassafras soil data.
| Nambsheim | Tama | Gross-Umstadt | Sassafras | Lleida | |
|---|---|---|---|---|---|
| fomc_path_2 | OK | OK | OK | C | OK |
| dfop_path_2 | OK | C | OK | C | OK |
| sforb_path_2 | OK | OK | OK | C | OK |
Using the two-component error model, all separate fits converge with the exception of the alternative pathway fit with DFOP used for the parent and the Sassafras dataset.
f_saem_2 <- mhmkin(list(f_sep_2_const, f_sep_2_tc),
no_random_effect = illparms(cyan_saem_full[2:4, ]),
cluster = cl_path_2)| const | tc | |
|---|---|---|
| fomc_path_2 | E | OK |
| dfop_path_2 | OK | OK |
| sforb_path_2 | OK | OK |
The hierarchical fits for the alternative pathway completed successfully, with the exception of the model using FOMC for the parent compound and constant variance as the error model.
| const | tc | |
|---|---|---|
| fomc_path_2 | E | sd(f_JSE76_qlogis) |
| dfop_path_2 | sd(f_JCZ38_qlogis), sd(f_JSE76_qlogis) | sd(f_JCZ38_qlogis), sd(f_JSE76_qlogis) |
| sforb_path_2 | sd(f_JCZ38_qlogis), sd(f_JSE76_qlogis) | sd(f_JCZ38_qlogis), sd(f_JSE76_qlogis) |
In all biphasic fits (DFOP or SFORB for the parent compound), the random effects for the formation fractions for the pathways from JCZ38 to JSE76, and for the reverse pathway from JSE76 to JCZ38 are ill-defined.
| npar | AIC | BIC | Lik | |
|---|---|---|---|---|
| fomc_path_2 tc | 21 | 2249.0 | 2240.8 | -1103.5 |
| dfop_path_2 tc | 22 | 2234.4 | 2225.8 | -1095.2 |
| sforb_path_2 tc | 22 | 2239.7 | 2231.1 | -1097.9 |
| npar | AIC | BIC | Lik | |
|---|---|---|---|---|
| dfop_path_2 const | 22 | 2288.4 | 2279.8 | -1122.2 |
| sforb_path_2 const | 22 | 2283.3 | 2274.7 | -1119.7 |
| dfop_path_2 tc | 22 | 2234.4 | 2225.8 | -1095.2 |
| sforb_path_2 tc | 22 | 2239.7 | 2231.1 | -1097.9 |
The variants using the biexponential models DFOP and SFORB for the parent compound and the two-component error model give the lowest AIC and BIC values and are plotted below. Compared with the original pathway, the AIC and BIC values indicate a large improvement. This is confirmed by the plots, which show that the metabolite JCZ38 is fitted much better with this model.
plot(f_saem_2[["fomc_path_2", "tc"]])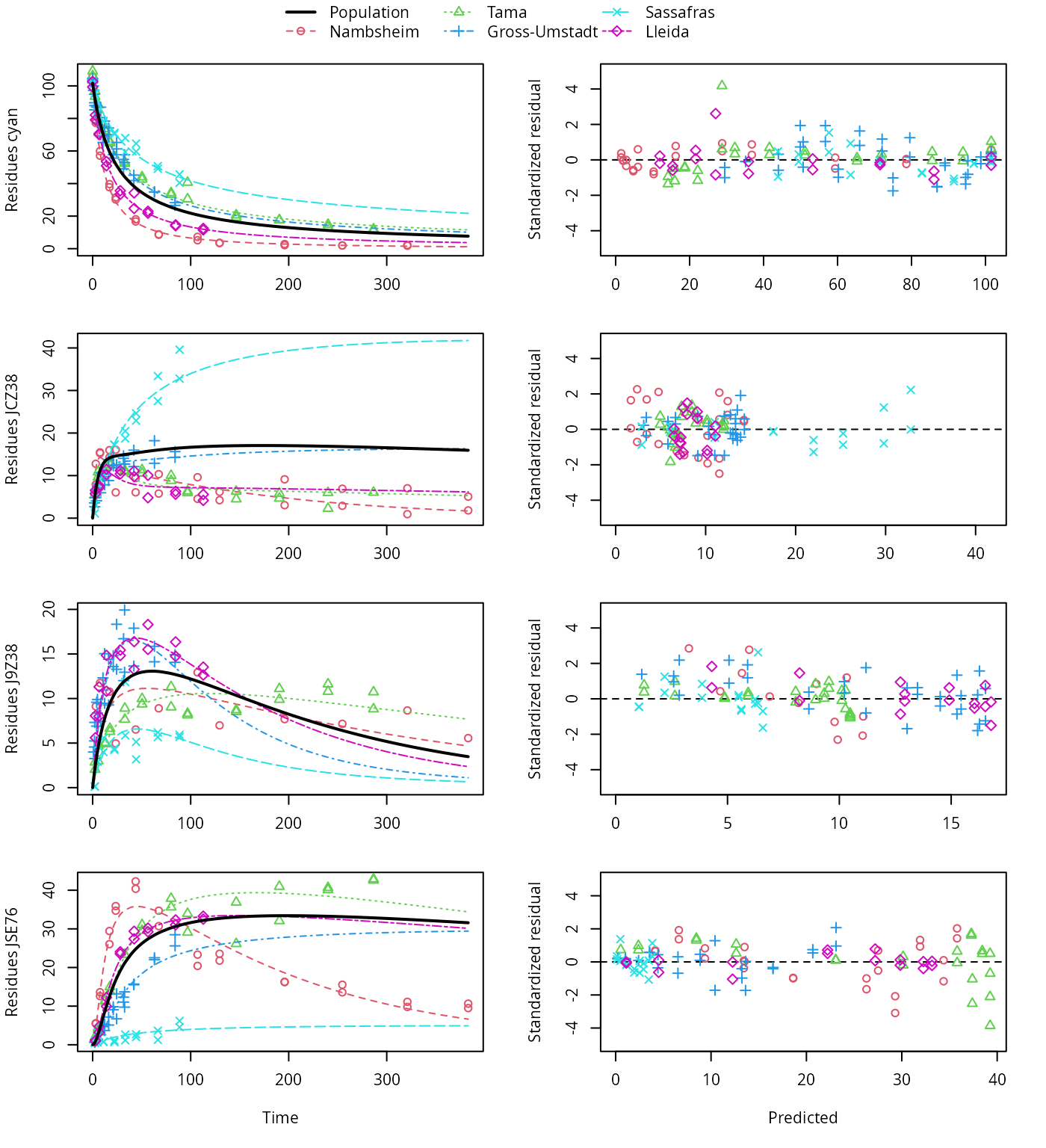
FOMC pathway fit with two-component error, alternative pathway
plot(f_saem_2[["dfop_path_2", "tc"]])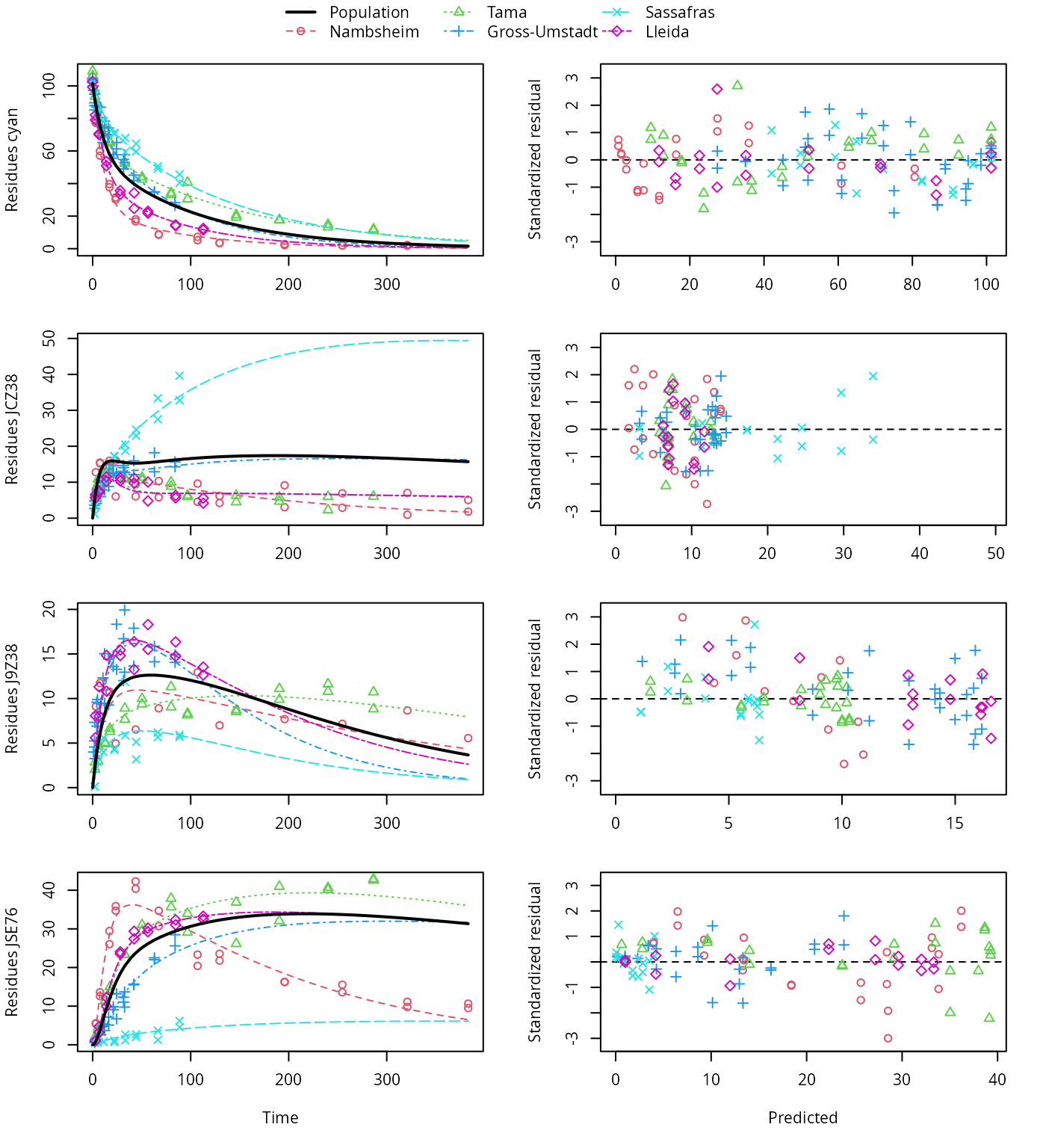
DFOP pathway fit with two-component error, alternative pathway
plot(f_saem_2[["sforb_path_2", "tc"]])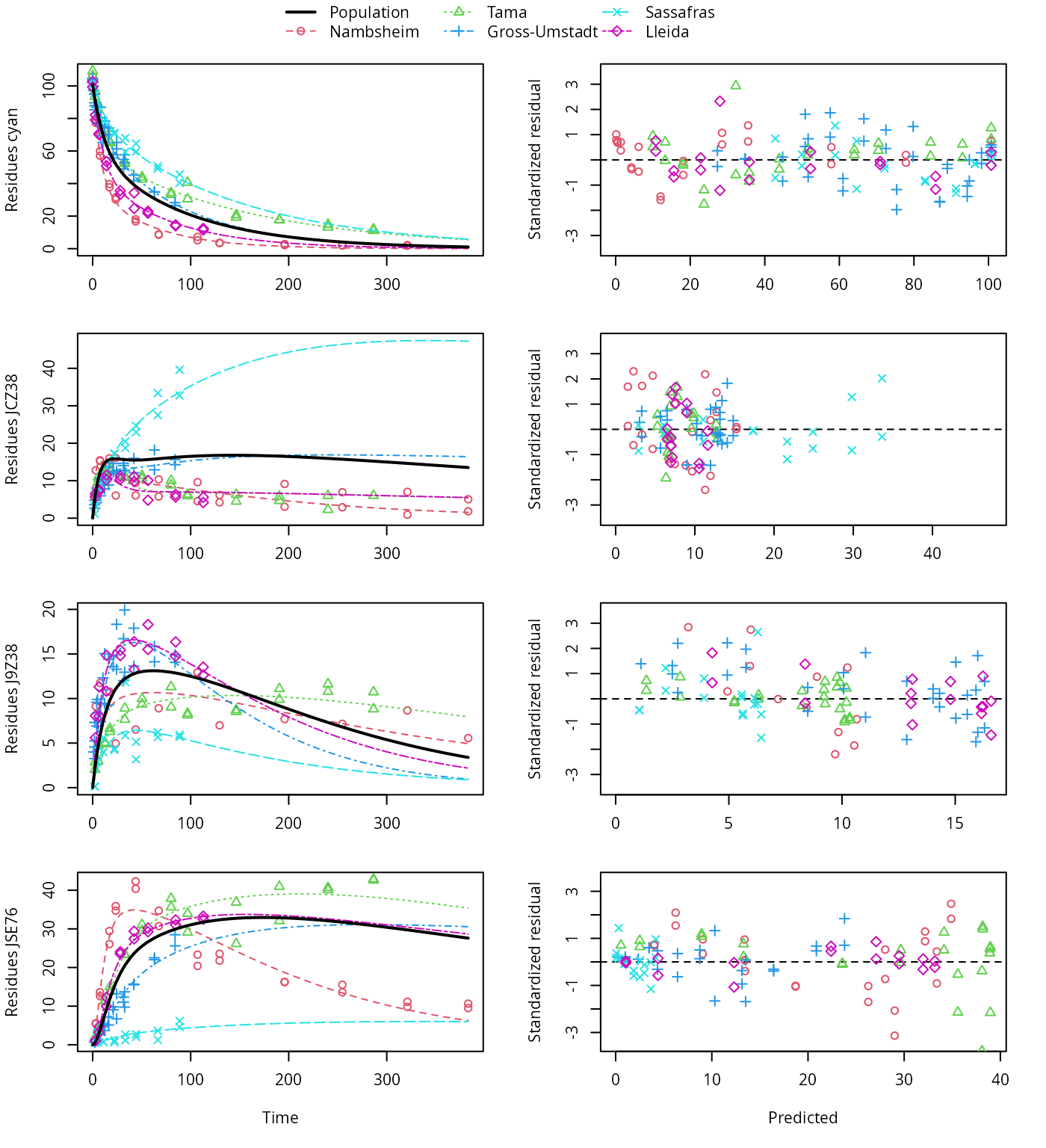
SFORB pathway fit with two-component error, alternative pathway
Refinement of alternative pathway fits
All ill-defined random effects that were identified in the parent
only fits and in the above pathway fits, are excluded for the final
evaluations below. For this purpose, a list of character vectors is
created below that can be indexed by row and column indices, and which
contains the degradation parameter names for which random effects should
be excluded for each of the hierarchical fits contained in
f_saem_2.
no_ranef <- matrix(list(), nrow = 3, ncol = 2, dimnames = dimnames(f_saem_2))
no_ranef[["fomc_path_2", "const"]] <- c("log_beta", "f_JCZ38_qlogis", "f_JSE76_qlogis")
no_ranef[["fomc_path_2", "tc"]] <- c("cyan_0", "f_JCZ38_qlogis", "f_JSE76_qlogis")
no_ranef[["dfop_path_2", "const"]] <- c("cyan_0", "f_JCZ38_qlogis", "f_JSE76_qlogis")
no_ranef[["dfop_path_2", "tc"]] <- c("cyan_0", "log_k1", "f_JCZ38_qlogis", "f_JSE76_qlogis")
no_ranef[["sforb_path_2", "const"]] <- c("cyan_free_0",
"f_JCZ38_qlogis", "f_JSE76_qlogis")
no_ranef[["sforb_path_2", "tc"]] <- c("cyan_free_0", "log_k_cyan_free_bound",
"f_JCZ38_qlogis", "f_JSE76_qlogis")
clusterExport(cl_path_2, "no_ranef")
f_saem_3 <- update(f_saem_2,
no_random_effect = no_ranef,
cluster = cl_path_2)| const | tc | |
|---|---|---|
| fomc_path_2 | E | Fth |
| dfop_path_2 | Fth | Fth |
| sforb_path_2 | Fth | Fth |
With the exception of the FOMC pathway fit with constant variance, all updated fits completed successfully. However, the Fisher Information Matrix for the fixed effects (Fth) could not be inverted, so no confidence intervals for the optimised parameters are available.
| const | tc | |
|---|---|---|
| fomc_path_2 | E | |
| dfop_path_2 | ||
| sforb_path_2 |
| npar | AIC | BIC | Lik | |
|---|---|---|---|---|
| fomc_path_2 tc | 19 | 2249.1 | 2241.6 | -1105.5 |
| dfop_path_2 tc | 20 | 2237.3 | 2229.5 | -1098.6 |
| sforb_path_2 tc | 20 | 2241.3 | 2233.5 | -1100.7 |
| npar | AIC | BIC | Lik | |
|---|---|---|---|---|
| dfop_path_2 const | 20 | 2282.2 | 2274.4 | -1121.1 |
| sforb_path_2 const | 20 | 2279.7 | 2271.9 | -1119.9 |
| dfop_path_2 tc | 20 | 2237.3 | 2229.5 | -1098.6 |
| sforb_path_2 tc | 20 | 2241.3 | 2233.5 | -1100.7 |
While the AIC and BIC values of the best fit (DFOP pathway fit with two-component error) are lower than in the previous fits with the alternative pathway, the practical value of these refined evaluations is limited as no confidence intervals are obtained.
stopCluster(cl_path_2)Conclusion
It was demonstrated that a relatively complex transformation pathway with parallel formation of two primary metabolites and one secondary metabolite can be fitted even if the data in the individual datasets are quite different and partly only cover the formation phase.
The run times of the pathway fits were several hours, limiting the practical feasibility of iterative refinements based on ill-defined parameters and of alternative checks of parameter identifiability based on multistart runs.
Acknowledgements
The helpful comments by Janina Wöltjen of the German Environment Agency are gratefully acknowledged.
Appendix
Plots of fits that were not refined further
plot(f_saem_1[["sfo_path_1", "tc"]])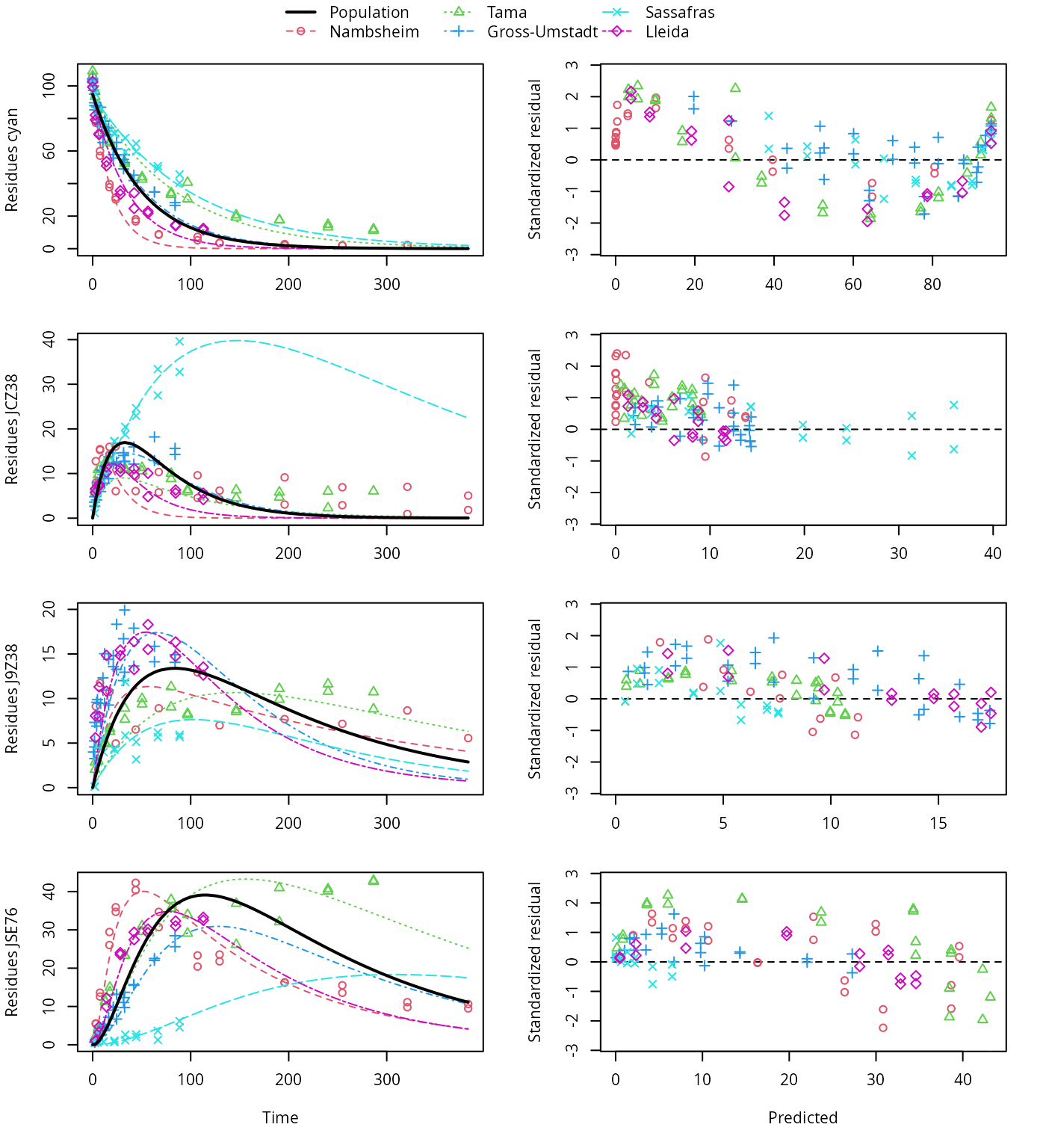
SFO pathway fit with two-component error
plot(f_saem_1[["fomc_path_1", "tc"]])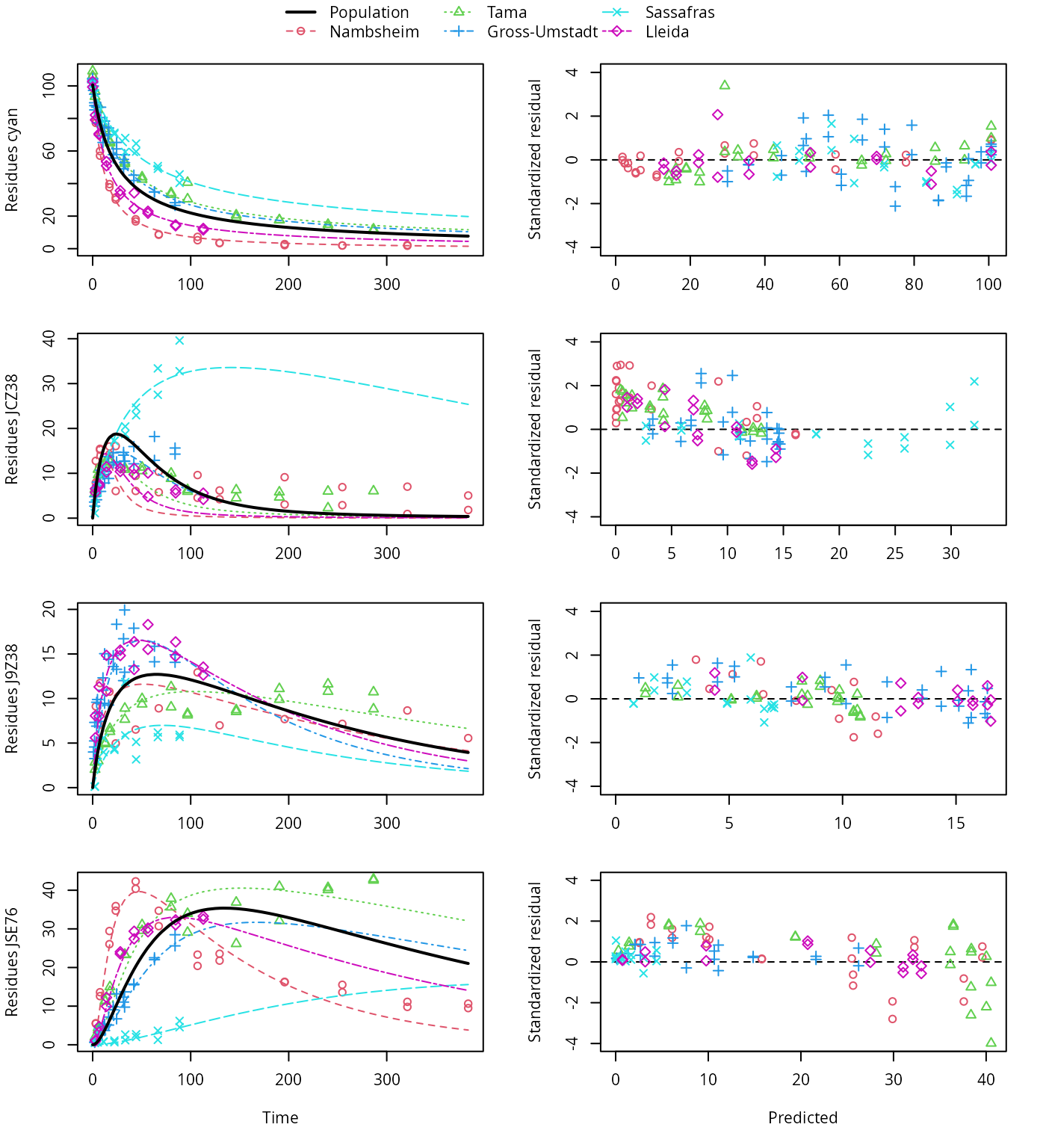
FOMC pathway fit with two-component error
plot(f_saem_1[["sforb_path_1", "tc"]])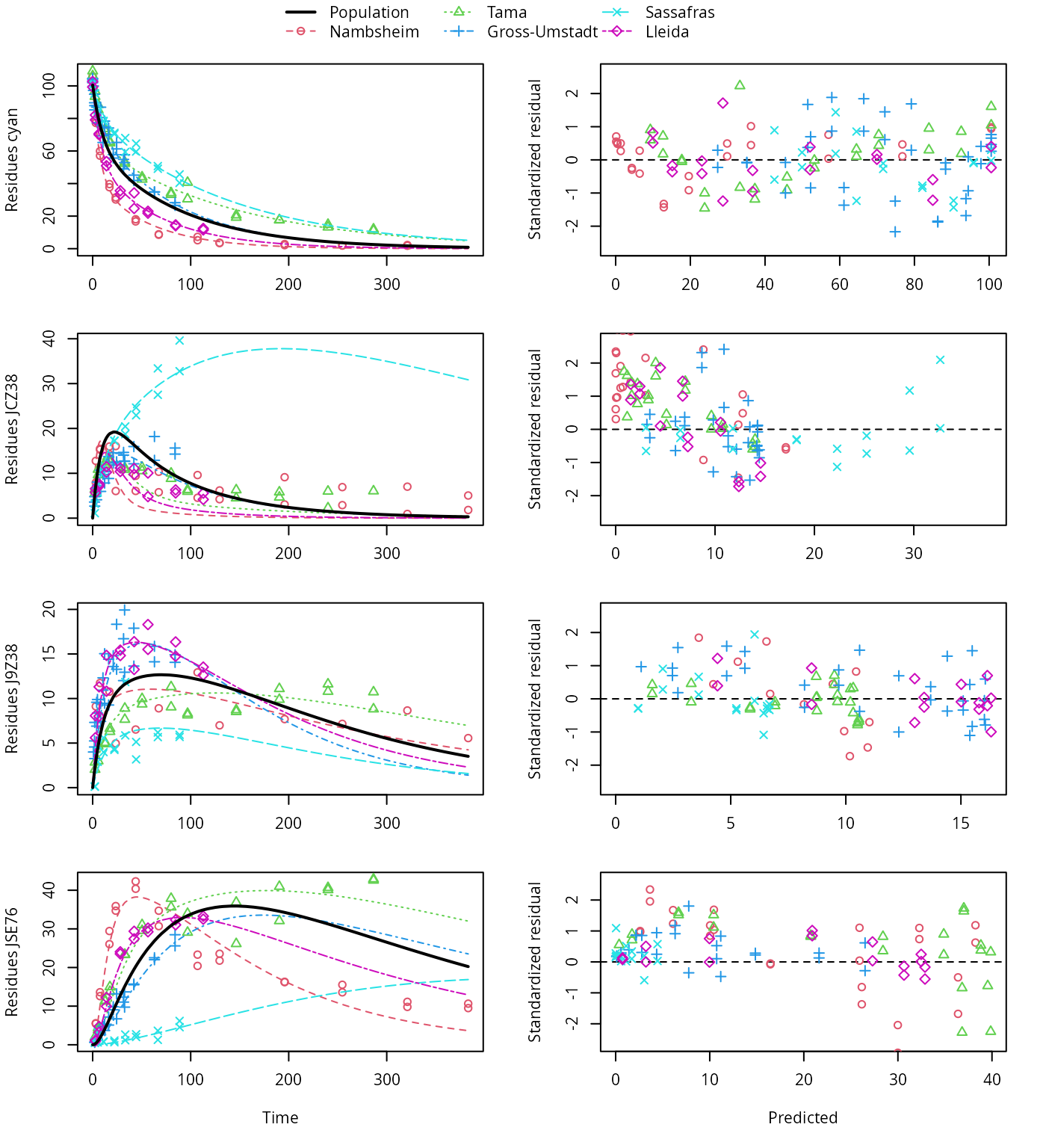
HS pathway fit with two-component error
Hierarchical fit listings
Pathway 1
saemix version used for fitting: 3.3
mkin version used for pre-fitting: 1.2.10
R version used for fitting: 4.5.0
Date of fit: Mon May 12 20:35:03 2025
Date of summary: Mon May 12 21:06:22 2025
Equations:
d_cyan/dt = - k_cyan * cyan
d_JCZ38/dt = + f_cyan_to_JCZ38 * k_cyan * cyan - k_JCZ38 * JCZ38
d_J9Z38/dt = + f_cyan_to_J9Z38 * k_cyan * cyan - k_J9Z38 * J9Z38
d_JSE76/dt = + f_JCZ38_to_JSE76 * k_JCZ38 * JCZ38 - k_JSE76 * JSE76
Data:
433 observations of 4 variable(s) grouped in 5 datasets
Model predictions using solution type deSolve
Fitted in 445.08 s
Using 300, 100 iterations and 10 chains
Variance model: Constant variance
Starting values for degradation parameters:
cyan_0 log_k_cyan log_k_JCZ38 log_k_J9Z38 log_k_JSE76
95.3304 -3.8459 -3.1305 -5.0678 -5.3196
f_cyan_ilr_1 f_cyan_ilr_2 f_JCZ38_qlogis
0.8158 23.5335 11.8774
Fixed degradation parameter values:
None
Starting values for random effects (square root of initial entries in omega):
cyan_0 log_k_cyan log_k_JCZ38 log_k_J9Z38 log_k_JSE76
cyan_0 4.797 0.0000 0.000 0.000 0.0000
log_k_cyan 0.000 0.9619 0.000 0.000 0.0000
log_k_JCZ38 0.000 0.0000 2.139 0.000 0.0000
log_k_J9Z38 0.000 0.0000 0.000 1.639 0.0000
log_k_JSE76 0.000 0.0000 0.000 0.000 0.7894
f_cyan_ilr_1 0.000 0.0000 0.000 0.000 0.0000
f_cyan_ilr_2 0.000 0.0000 0.000 0.000 0.0000
f_JCZ38_qlogis 0.000 0.0000 0.000 0.000 0.0000
f_cyan_ilr_1 f_cyan_ilr_2 f_JCZ38_qlogis
cyan_0 0.0000 0.000 0.00
log_k_cyan 0.0000 0.000 0.00
log_k_JCZ38 0.0000 0.000 0.00
log_k_J9Z38 0.0000 0.000 0.00
log_k_JSE76 0.0000 0.000 0.00
f_cyan_ilr_1 0.7714 0.000 0.00
f_cyan_ilr_2 0.0000 9.247 0.00
f_JCZ38_qlogis 0.0000 0.000 16.61
Starting values for error model parameters:
a.1
1
Results:
Likelihood computed by importance sampling
AIC BIC logLik
2693 2687 -1331
Optimised parameters:
est. lower upper
cyan_0 95.1279 9.354e+01 9.671e+01
log_k_cyan -3.8527 -4.367e+00 -3.338e+00
log_k_JCZ38 -3.0381 -4.187e+00 -1.889e+00
log_k_J9Z38 -5.0095 -5.623e+00 -4.396e+00
log_k_JSE76 -5.3357 -6.025e+00 -4.646e+00
f_cyan_ilr_1 0.8050 5.174e-01 1.093e+00
f_cyan_ilr_2 12.4820 -1.050e+06 1.051e+06
f_JCZ38_qlogis 1.2912 3.561e-01 2.226e+00
a.1 4.8393 NA NA
SD.log_k_cyan 0.5840 NA NA
SD.log_k_JCZ38 1.2740 NA NA
SD.log_k_J9Z38 0.3172 NA NA
SD.log_k_JSE76 0.5677 NA NA
SD.f_cyan_ilr_1 0.2623 NA NA
SD.f_cyan_ilr_2 1.3724 NA NA
SD.f_JCZ38_qlogis 0.1464 NA NA
Correlation is not available
Random effects:
est. lower upper
SD.log_k_cyan 0.5840 NA NA
SD.log_k_JCZ38 1.2740 NA NA
SD.log_k_J9Z38 0.3172 NA NA
SD.log_k_JSE76 0.5677 NA NA
SD.f_cyan_ilr_1 0.2623 NA NA
SD.f_cyan_ilr_2 1.3724 NA NA
SD.f_JCZ38_qlogis 0.1464 NA NA
Variance model:
est. lower upper
a.1 4.839 NA NA
Backtransformed parameters:
est. lower upper
cyan_0 95.127935 93.542456 96.713413
k_cyan 0.021221 0.012687 0.035497
k_JCZ38 0.047924 0.015189 0.151213
k_J9Z38 0.006674 0.003612 0.012332
k_JSE76 0.004817 0.002417 0.009601
f_cyan_to_JCZ38 0.757402 NA NA
f_cyan_to_J9Z38 0.242597 NA NA
f_JCZ38_to_JSE76 0.784347 0.588098 0.902582
Resulting formation fractions:
ff
cyan_JCZ38 7.574e-01
cyan_J9Z38 2.426e-01
cyan_sink 9.839e-08
JCZ38_JSE76 7.843e-01
JCZ38_sink 2.157e-01
Estimated disappearance times:
DT50 DT90
cyan 32.66 108.50
JCZ38 14.46 48.05
J9Z38 103.86 345.00
JSE76 143.91 478.04
saemix version used for fitting: 3.3
mkin version used for pre-fitting: 1.2.10
R version used for fitting: 4.5.0
Date of fit: Mon May 12 20:34:47 2025
Date of summary: Mon May 12 21:06:22 2025
Equations:
d_cyan/dt = - k_cyan * cyan
d_JCZ38/dt = + f_cyan_to_JCZ38 * k_cyan * cyan - k_JCZ38 * JCZ38
d_J9Z38/dt = + f_cyan_to_J9Z38 * k_cyan * cyan - k_J9Z38 * J9Z38
d_JSE76/dt = + f_JCZ38_to_JSE76 * k_JCZ38 * JCZ38 - k_JSE76 * JSE76
Data:
433 observations of 4 variable(s) grouped in 5 datasets
Model predictions using solution type deSolve
Fitted in 428.46 s
Using 300, 100 iterations and 10 chains
Variance model: Two-component variance function
Starting values for degradation parameters:
cyan_0 log_k_cyan log_k_JCZ38 log_k_J9Z38 log_k_JSE76
96.0039 -3.8907 -3.1276 -5.0069 -4.9367
f_cyan_ilr_1 f_cyan_ilr_2 f_JCZ38_qlogis
0.7937 22.3422 17.8932
Fixed degradation parameter values:
None
Starting values for random effects (square root of initial entries in omega):
cyan_0 log_k_cyan log_k_JCZ38 log_k_J9Z38 log_k_JSE76
cyan_0 4.859 0.000 0.00 0.00 0.0000
log_k_cyan 0.000 0.962 0.00 0.00 0.0000
log_k_JCZ38 0.000 0.000 2.04 0.00 0.0000
log_k_J9Z38 0.000 0.000 0.00 1.72 0.0000
log_k_JSE76 0.000 0.000 0.00 0.00 0.9076
f_cyan_ilr_1 0.000 0.000 0.00 0.00 0.0000
f_cyan_ilr_2 0.000 0.000 0.00 0.00 0.0000
f_JCZ38_qlogis 0.000 0.000 0.00 0.00 0.0000
f_cyan_ilr_1 f_cyan_ilr_2 f_JCZ38_qlogis
cyan_0 0.0000 0.000 0.00
log_k_cyan 0.0000 0.000 0.00
log_k_JCZ38 0.0000 0.000 0.00
log_k_J9Z38 0.0000 0.000 0.00
log_k_JSE76 0.0000 0.000 0.00
f_cyan_ilr_1 0.7598 0.000 0.00
f_cyan_ilr_2 0.0000 8.939 0.00
f_JCZ38_qlogis 0.0000 0.000 14.49
Starting values for error model parameters:
a.1 b.1
1 1
Results:
Likelihood computed by importance sampling
AIC BIC logLik
2658 2651 -1312
Optimised parameters:
est. lower upper
cyan_0 94.81681 NA NA
log_k_cyan -3.91558 NA NA
log_k_JCZ38 -3.12715 NA NA
log_k_J9Z38 -5.04840 NA NA
log_k_JSE76 -5.10443 NA NA
f_cyan_ilr_1 0.80760 NA NA
f_cyan_ilr_2 48.66960 NA NA
f_JCZ38_qlogis 3.03397 NA NA
a.1 3.93879 NA NA
b.1 0.08057 NA NA
SD.log_k_cyan 0.58921 NA NA
SD.log_k_JCZ38 1.29813 NA NA
SD.log_k_J9Z38 0.68372 NA NA
SD.log_k_JSE76 0.35128 NA NA
SD.f_cyan_ilr_1 0.38352 NA NA
SD.f_cyan_ilr_2 4.98884 NA NA
SD.f_JCZ38_qlogis 1.75636 NA NA
Correlation is not available
Random effects:
est. lower upper
SD.log_k_cyan 0.5892 NA NA
SD.log_k_JCZ38 1.2981 NA NA
SD.log_k_J9Z38 0.6837 NA NA
SD.log_k_JSE76 0.3513 NA NA
SD.f_cyan_ilr_1 0.3835 NA NA
SD.f_cyan_ilr_2 4.9888 NA NA
SD.f_JCZ38_qlogis 1.7564 NA NA
Variance model:
est. lower upper
a.1 3.93879 NA NA
b.1 0.08057 NA NA
Backtransformed parameters:
est. lower upper
cyan_0 94.81681 NA NA
k_cyan 0.01993 NA NA
k_JCZ38 0.04384 NA NA
k_J9Z38 0.00642 NA NA
k_JSE76 0.00607 NA NA
f_cyan_to_JCZ38 0.75807 NA NA
f_cyan_to_J9Z38 0.24193 NA NA
f_JCZ38_to_JSE76 0.95409 NA NA
Resulting formation fractions:
ff
cyan_JCZ38 0.75807
cyan_J9Z38 0.24193
cyan_sink 0.00000
JCZ38_JSE76 0.95409
JCZ38_sink 0.04591
Estimated disappearance times:
DT50 DT90
cyan 34.78 115.54
JCZ38 15.81 52.52
J9Z38 107.97 358.68
JSE76 114.20 379.35
saemix version used for fitting: 3.3
mkin version used for pre-fitting: 1.2.10
R version used for fitting: 4.5.0
Date of fit: Mon May 12 20:35:21 2025
Date of summary: Mon May 12 21:06:22 2025
Equations:
d_cyan/dt = - (alpha/beta) * 1/((time/beta) + 1) * cyan
d_JCZ38/dt = + f_cyan_to_JCZ38 * (alpha/beta) * 1/((time/beta) + 1) *
cyan - k_JCZ38 * JCZ38
d_J9Z38/dt = + f_cyan_to_J9Z38 * (alpha/beta) * 1/((time/beta) + 1) *
cyan - k_J9Z38 * J9Z38
d_JSE76/dt = + f_JCZ38_to_JSE76 * k_JCZ38 * JCZ38 - k_JSE76 * JSE76
Data:
433 observations of 4 variable(s) grouped in 5 datasets
Model predictions using solution type deSolve
Fitted in 462.739 s
Using 300, 100 iterations and 10 chains
Variance model: Constant variance
Starting values for degradation parameters:
cyan_0 log_k_JCZ38 log_k_J9Z38 log_k_JSE76 f_cyan_ilr_1
101.2314 -3.3680 -5.1108 -5.9416 0.7144
f_cyan_ilr_2 f_JCZ38_qlogis log_alpha log_beta
7.0229 14.9234 -0.1791 2.9811
Fixed degradation parameter values:
None
Starting values for random effects (square root of initial entries in omega):
cyan_0 log_k_JCZ38 log_k_J9Z38 log_k_JSE76 f_cyan_ilr_1
cyan_0 5.416 0.000 0.0 0.000 0.0000
log_k_JCZ38 0.000 2.439 0.0 0.000 0.0000
log_k_J9Z38 0.000 0.000 1.7 0.000 0.0000
log_k_JSE76 0.000 0.000 0.0 1.856 0.0000
f_cyan_ilr_1 0.000 0.000 0.0 0.000 0.7164
f_cyan_ilr_2 0.000 0.000 0.0 0.000 0.0000
f_JCZ38_qlogis 0.000 0.000 0.0 0.000 0.0000
log_alpha 0.000 0.000 0.0 0.000 0.0000
log_beta 0.000 0.000 0.0 0.000 0.0000
f_cyan_ilr_2 f_JCZ38_qlogis log_alpha log_beta
cyan_0 0.00 0.00 0.0000 0.0000
log_k_JCZ38 0.00 0.00 0.0000 0.0000
log_k_J9Z38 0.00 0.00 0.0000 0.0000
log_k_JSE76 0.00 0.00 0.0000 0.0000
f_cyan_ilr_1 0.00 0.00 0.0000 0.0000
f_cyan_ilr_2 11.57 0.00 0.0000 0.0000
f_JCZ38_qlogis 0.00 18.81 0.0000 0.0000
log_alpha 0.00 0.00 0.4144 0.0000
log_beta 0.00 0.00 0.0000 0.5077
Starting values for error model parameters:
a.1
1
Results:
Likelihood computed by importance sampling
AIC BIC logLik
2428 2421 -1196
Optimised parameters:
est. lower upper
cyan_0 101.1664 98.51265 103.8202
log_k_JCZ38 -3.3883 -4.78250 -1.9941
log_k_J9Z38 -5.3087 -5.91564 -4.7017
log_k_JSE76 -6.1313 -7.30061 -4.9619
f_cyan_ilr_1 0.7456 0.43782 1.0534
f_cyan_ilr_2 0.8181 0.24956 1.3866
f_JCZ38_qlogis 2.0467 0.61165 3.4817
log_alpha -0.2391 -0.62806 0.1499
log_beta 2.8739 2.67664 3.0711
a.1 3.4160 3.17960 3.6525
SD.cyan_0 2.4355 0.40399 4.4671
SD.log_k_JCZ38 1.5654 0.57311 2.5576
SD.log_k_J9Z38 0.4645 -0.06533 0.9943
SD.log_k_JSE76 0.9841 0.10738 1.8609
SD.f_cyan_ilr_1 0.3285 0.10546 0.5515
SD.f_cyan_ilr_2 0.2276 -0.38711 0.8424
SD.f_JCZ38_qlogis 0.8340 -0.20970 1.8777
SD.log_alpha 0.4250 0.16017 0.6898
Correlation:
cyan_0 l__JCZ3 l__J9Z3 l__JSE7 f_cy__1 f_cy__2 f_JCZ38 log_lph
log_k_JCZ38 -0.0159
log_k_J9Z38 -0.0546 0.0080
log_k_JSE76 -0.0337 0.0016 0.0074
f_cyan_ilr_1 -0.0095 0.0194 -0.1573 0.0003
f_cyan_ilr_2 -0.2733 0.0799 0.3059 0.0263 0.0125
f_JCZ38_qlogis 0.0755 -0.0783 -0.0516 0.1222 -0.1155 -0.5231
log_alpha -0.0567 0.0120 0.0351 0.0189 0.0040 0.0829 -0.0502
log_beta -0.2980 0.0461 0.1382 0.0758 0.0209 0.4079 -0.2053 0.2759
Random effects:
est. lower upper
SD.cyan_0 2.4355 0.40399 4.4671
SD.log_k_JCZ38 1.5654 0.57311 2.5576
SD.log_k_J9Z38 0.4645 -0.06533 0.9943
SD.log_k_JSE76 0.9841 0.10738 1.8609
SD.f_cyan_ilr_1 0.3285 0.10546 0.5515
SD.f_cyan_ilr_2 0.2276 -0.38711 0.8424
SD.f_JCZ38_qlogis 0.8340 -0.20970 1.8777
SD.log_alpha 0.4250 0.16017 0.6898
Variance model:
est. lower upper
a.1 3.416 3.18 3.652
Backtransformed parameters:
est. lower upper
cyan_0 1.012e+02 9.851e+01 103.82023
k_JCZ38 3.377e-02 8.375e-03 0.13614
k_J9Z38 4.948e-03 2.697e-03 0.00908
k_JSE76 2.174e-03 6.751e-04 0.00700
f_cyan_to_JCZ38 6.389e-01 NA NA
f_cyan_to_J9Z38 2.226e-01 NA NA
f_JCZ38_to_JSE76 8.856e-01 6.483e-01 0.97016
alpha 7.873e-01 5.336e-01 1.16166
beta 1.771e+01 1.454e+01 21.56509
Resulting formation fractions:
ff
cyan_JCZ38 0.6389
cyan_J9Z38 0.2226
cyan_sink 0.1385
JCZ38_JSE76 0.8856
JCZ38_sink 0.1144
Estimated disappearance times:
DT50 DT90 DT50back
cyan 25.00 312.06 93.94
JCZ38 20.53 68.19 NA
J9Z38 140.07 465.32 NA
JSE76 318.86 1059.22 NA
saemix version used for fitting: 3.3
mkin version used for pre-fitting: 1.2.10
R version used for fitting: 4.5.0
Date of fit: Mon May 12 20:35:31 2025
Date of summary: Mon May 12 21:06:22 2025
Equations:
d_cyan/dt = - (alpha/beta) * 1/((time/beta) + 1) * cyan
d_JCZ38/dt = + f_cyan_to_JCZ38 * (alpha/beta) * 1/((time/beta) + 1) *
cyan - k_JCZ38 * JCZ38
d_J9Z38/dt = + f_cyan_to_J9Z38 * (alpha/beta) * 1/((time/beta) + 1) *
cyan - k_J9Z38 * J9Z38
d_JSE76/dt = + f_JCZ38_to_JSE76 * k_JCZ38 * JCZ38 - k_JSE76 * JSE76
Data:
433 observations of 4 variable(s) grouped in 5 datasets
Model predictions using solution type deSolve
Fitted in 472.067 s
Using 300, 100 iterations and 10 chains
Variance model: Two-component variance function
Starting values for degradation parameters:
cyan_0 log_k_JCZ38 log_k_J9Z38 log_k_JSE76 f_cyan_ilr_1
101.13294 -3.32499 -5.09097 -5.93566 0.71359
f_cyan_ilr_2 f_JCZ38_qlogis log_alpha log_beta
10.30315 14.62272 -0.09633 3.10634
Fixed degradation parameter values:
None
Starting values for random effects (square root of initial entries in omega):
cyan_0 log_k_JCZ38 log_k_J9Z38 log_k_JSE76 f_cyan_ilr_1
cyan_0 5.649 0.000 0.00 0.00 0.0000
log_k_JCZ38 0.000 2.319 0.00 0.00 0.0000
log_k_J9Z38 0.000 0.000 1.73 0.00 0.0000
log_k_JSE76 0.000 0.000 0.00 1.86 0.0000
f_cyan_ilr_1 0.000 0.000 0.00 0.00 0.7183
f_cyan_ilr_2 0.000 0.000 0.00 0.00 0.0000
f_JCZ38_qlogis 0.000 0.000 0.00 0.00 0.0000
log_alpha 0.000 0.000 0.00 0.00 0.0000
log_beta 0.000 0.000 0.00 0.00 0.0000
f_cyan_ilr_2 f_JCZ38_qlogis log_alpha log_beta
cyan_0 0.00 0.00 0.0000 0.0000
log_k_JCZ38 0.00 0.00 0.0000 0.0000
log_k_J9Z38 0.00 0.00 0.0000 0.0000
log_k_JSE76 0.00 0.00 0.0000 0.0000
f_cyan_ilr_1 0.00 0.00 0.0000 0.0000
f_cyan_ilr_2 12.85 0.00 0.0000 0.0000
f_JCZ38_qlogis 0.00 18.54 0.0000 0.0000
log_alpha 0.00 0.00 0.3142 0.0000
log_beta 0.00 0.00 0.0000 0.7333
Starting values for error model parameters:
a.1 b.1
1 1
Results:
Likelihood computed by importance sampling
AIC BIC logLik
2424 2416 -1193
Optimised parameters:
est. lower upper
cyan_0 100.65667 NA NA
log_k_JCZ38 -3.45782 NA NA
log_k_J9Z38 -5.23476 NA NA
log_k_JSE76 -5.71827 NA NA
f_cyan_ilr_1 0.68389 NA NA
f_cyan_ilr_2 0.61027 NA NA
f_JCZ38_qlogis 116.27482 NA NA
log_alpha -0.14484 NA NA
log_beta 3.03220 NA NA
a.1 3.11051 NA NA
b.1 0.04508 NA NA
SD.log_k_JCZ38 1.39961 NA NA
SD.log_k_J9Z38 0.57920 NA NA
SD.log_k_JSE76 0.68364 NA NA
SD.f_cyan_ilr_1 0.31477 NA NA
SD.f_cyan_ilr_2 0.37716 NA NA
SD.f_JCZ38_qlogis 5.52695 NA NA
SD.log_alpha 0.22823 NA NA
SD.log_beta 0.39161 NA NA
Correlation is not available
Random effects:
est. lower upper
SD.log_k_JCZ38 1.3996 NA NA
SD.log_k_J9Z38 0.5792 NA NA
SD.log_k_JSE76 0.6836 NA NA
SD.f_cyan_ilr_1 0.3148 NA NA
SD.f_cyan_ilr_2 0.3772 NA NA
SD.f_JCZ38_qlogis 5.5270 NA NA
SD.log_alpha 0.2282 NA NA
SD.log_beta 0.3916 NA NA
Variance model:
est. lower upper
a.1 3.11051 NA NA
b.1 0.04508 NA NA
Backtransformed parameters:
est. lower upper
cyan_0 1.007e+02 NA NA
k_JCZ38 3.150e-02 NA NA
k_J9Z38 5.328e-03 NA NA
k_JSE76 3.285e-03 NA NA
f_cyan_to_JCZ38 5.980e-01 NA NA
f_cyan_to_J9Z38 2.273e-01 NA NA
f_JCZ38_to_JSE76 1.000e+00 NA NA
alpha 8.652e-01 NA NA
beta 2.074e+01 NA NA
Resulting formation fractions:
ff
cyan_JCZ38 0.5980
cyan_J9Z38 0.2273
cyan_sink 0.1746
JCZ38_JSE76 1.0000
JCZ38_sink 0.0000
Estimated disappearance times:
DT50 DT90 DT50back
cyan 25.48 276.2 83.15
JCZ38 22.01 73.1 NA
J9Z38 130.09 432.2 NA
JSE76 210.98 700.9 NA
saemix version used for fitting: 3.3
mkin version used for pre-fitting: 1.2.10
R version used for fitting: 4.5.0
Date of fit: Mon May 12 20:36:02 2025
Date of summary: Mon May 12 21:06:22 2025
Equations:
d_cyan/dt = - ((k1 * g * exp(-k1 * time) + k2 * (1 - g) * exp(-k2 *
time)) / (g * exp(-k1 * time) + (1 - g) * exp(-k2 * time)))
* cyan
d_JCZ38/dt = + f_cyan_to_JCZ38 * ((k1 * g * exp(-k1 * time) + k2 * (1 -
g) * exp(-k2 * time)) / (g * exp(-k1 * time) + (1 - g) *
exp(-k2 * time))) * cyan - k_JCZ38 * JCZ38
d_J9Z38/dt = + f_cyan_to_J9Z38 * ((k1 * g * exp(-k1 * time) + k2 * (1 -
g) * exp(-k2 * time)) / (g * exp(-k1 * time) + (1 - g) *
exp(-k2 * time))) * cyan - k_J9Z38 * J9Z38
d_JSE76/dt = + f_JCZ38_to_JSE76 * k_JCZ38 * JCZ38 - k_JSE76 * JSE76
Data:
433 observations of 4 variable(s) grouped in 5 datasets
Model predictions using solution type deSolve
Fitted in 503.671 s
Using 300, 100 iterations and 10 chains
Variance model: Constant variance
Starting values for degradation parameters:
cyan_0 log_k_JCZ38 log_k_J9Z38 log_k_JSE76 f_cyan_ilr_1
102.0643 -3.4008 -5.0024 -5.8612 0.6855
f_cyan_ilr_2 f_JCZ38_qlogis log_k1 log_k2 g_qlogis
1.2366 13.6901 -1.8641 -4.5063 -0.6468
Fixed degradation parameter values:
None
Starting values for random effects (square root of initial entries in omega):
cyan_0 log_k_JCZ38 log_k_J9Z38 log_k_JSE76 f_cyan_ilr_1
cyan_0 4.466 0.000 0.000 0.000 0.0000
log_k_JCZ38 0.000 2.382 0.000 0.000 0.0000
log_k_J9Z38 0.000 0.000 1.595 0.000 0.0000
log_k_JSE76 0.000 0.000 0.000 1.245 0.0000
f_cyan_ilr_1 0.000 0.000 0.000 0.000 0.6852
f_cyan_ilr_2 0.000 0.000 0.000 0.000 0.0000
f_JCZ38_qlogis 0.000 0.000 0.000 0.000 0.0000
log_k1 0.000 0.000 0.000 0.000 0.0000
log_k2 0.000 0.000 0.000 0.000 0.0000
g_qlogis 0.000 0.000 0.000 0.000 0.0000
f_cyan_ilr_2 f_JCZ38_qlogis log_k1 log_k2 g_qlogis
cyan_0 0.00 0.00 0.0000 0.0000 0.000
log_k_JCZ38 0.00 0.00 0.0000 0.0000 0.000
log_k_J9Z38 0.00 0.00 0.0000 0.0000 0.000
log_k_JSE76 0.00 0.00 0.0000 0.0000 0.000
f_cyan_ilr_1 0.00 0.00 0.0000 0.0000 0.000
f_cyan_ilr_2 1.28 0.00 0.0000 0.0000 0.000
f_JCZ38_qlogis 0.00 16.08 0.0000 0.0000 0.000
log_k1 0.00 0.00 0.9866 0.0000 0.000
log_k2 0.00 0.00 0.0000 0.5953 0.000
g_qlogis 0.00 0.00 0.0000 0.0000 1.583
Starting values for error model parameters:
a.1
1
Results:
Likelihood computed by importance sampling
AIC BIC logLik
2403 2395 -1182
Optimised parameters:
est. lower upper
cyan_0 102.5565 NA NA
log_k_JCZ38 -3.4729 NA NA
log_k_J9Z38 -5.1533 NA NA
log_k_JSE76 -5.6669 NA NA
f_cyan_ilr_1 0.6665 NA NA
f_cyan_ilr_2 0.5191 NA NA
f_JCZ38_qlogis 37.0113 NA NA
log_k1 -1.8497 NA NA
log_k2 -4.4931 NA NA
g_qlogis -0.6383 NA NA
a.1 3.2397 NA NA
SD.log_k_JCZ38 1.4286 NA NA
SD.log_k_J9Z38 0.5312 NA NA
SD.log_k_JSE76 0.6627 NA NA
SD.f_cyan_ilr_1 0.3013 NA NA
SD.f_cyan_ilr_2 0.2980 NA NA
SD.f_JCZ38_qlogis 0.1637 NA NA
SD.log_k1 0.5069 NA NA
SD.log_k2 0.3828 NA NA
SD.g_qlogis 0.8641 NA NA
Correlation is not available
Random effects:
est. lower upper
SD.log_k_JCZ38 1.4286 NA NA
SD.log_k_J9Z38 0.5312 NA NA
SD.log_k_JSE76 0.6627 NA NA
SD.f_cyan_ilr_1 0.3013 NA NA
SD.f_cyan_ilr_2 0.2980 NA NA
SD.f_JCZ38_qlogis 0.1637 NA NA
SD.log_k1 0.5069 NA NA
SD.log_k2 0.3828 NA NA
SD.g_qlogis 0.8641 NA NA
Variance model:
est. lower upper
a.1 3.24 NA NA
Backtransformed parameters:
est. lower upper
cyan_0 1.026e+02 NA NA
k_JCZ38 3.103e-02 NA NA
k_J9Z38 5.780e-03 NA NA
k_JSE76 3.459e-03 NA NA
f_cyan_to_JCZ38 5.813e-01 NA NA
f_cyan_to_J9Z38 2.265e-01 NA NA
f_JCZ38_to_JSE76 1.000e+00 NA NA
k1 1.573e-01 NA NA
k2 1.119e-02 NA NA
g 3.456e-01 NA NA
Resulting formation fractions:
ff
cyan_JCZ38 0.5813
cyan_J9Z38 0.2265
cyan_sink 0.1922
JCZ38_JSE76 1.0000
JCZ38_sink 0.0000
Estimated disappearance times:
DT50 DT90 DT50back DT50_k1 DT50_k2
cyan 25.23 167.94 50.55 4.407 61.97
JCZ38 22.34 74.22 NA NA NA
J9Z38 119.92 398.36 NA NA NA
JSE76 200.41 665.76 NA NA NA
saemix version used for fitting: 3.3
mkin version used for pre-fitting: 1.2.10
R version used for fitting: 4.5.0
Date of fit: Mon May 12 20:38:54 2025
Date of summary: Mon May 12 21:06:22 2025
Equations:
d_cyan/dt = - ((k1 * g * exp(-k1 * time) + k2 * (1 - g) * exp(-k2 *
time)) / (g * exp(-k1 * time) + (1 - g) * exp(-k2 * time)))
* cyan
d_JCZ38/dt = + f_cyan_to_JCZ38 * ((k1 * g * exp(-k1 * time) + k2 * (1 -
g) * exp(-k2 * time)) / (g * exp(-k1 * time) + (1 - g) *
exp(-k2 * time))) * cyan - k_JCZ38 * JCZ38
d_J9Z38/dt = + f_cyan_to_J9Z38 * ((k1 * g * exp(-k1 * time) + k2 * (1 -
g) * exp(-k2 * time)) / (g * exp(-k1 * time) + (1 - g) *
exp(-k2 * time))) * cyan - k_J9Z38 * J9Z38
d_JSE76/dt = + f_JCZ38_to_JSE76 * k_JCZ38 * JCZ38 - k_JSE76 * JSE76
Data:
433 observations of 4 variable(s) grouped in 5 datasets
Model predictions using solution type deSolve
Fitted in 674.81 s
Using 300, 100 iterations and 10 chains
Variance model: Two-component variance function
Starting values for degradation parameters:
cyan_0 log_k_JCZ38 log_k_J9Z38 log_k_JSE76 f_cyan_ilr_1
101.3964 -3.3626 -4.9792 -5.8727 0.6814
f_cyan_ilr_2 f_JCZ38_qlogis log_k1 log_k2 g_qlogis
6.8713 13.6901 -1.9222 -4.5035 -0.7172
Fixed degradation parameter values:
None
Starting values for random effects (square root of initial entries in omega):
cyan_0 log_k_JCZ38 log_k_J9Z38 log_k_JSE76 f_cyan_ilr_1
cyan_0 5.317 0.000 0.000 0.000 0.0000
log_k_JCZ38 0.000 2.272 0.000 0.000 0.0000
log_k_J9Z38 0.000 0.000 1.633 0.000 0.0000
log_k_JSE76 0.000 0.000 0.000 1.271 0.0000
f_cyan_ilr_1 0.000 0.000 0.000 0.000 0.6839
f_cyan_ilr_2 0.000 0.000 0.000 0.000 0.0000
f_JCZ38_qlogis 0.000 0.000 0.000 0.000 0.0000
log_k1 0.000 0.000 0.000 0.000 0.0000
log_k2 0.000 0.000 0.000 0.000 0.0000
g_qlogis 0.000 0.000 0.000 0.000 0.0000
f_cyan_ilr_2 f_JCZ38_qlogis log_k1 log_k2 g_qlogis
cyan_0 0.00 0.00 0.0000 0.0000 0.000
log_k_JCZ38 0.00 0.00 0.0000 0.0000 0.000
log_k_J9Z38 0.00 0.00 0.0000 0.0000 0.000
log_k_JSE76 0.00 0.00 0.0000 0.0000 0.000
f_cyan_ilr_1 0.00 0.00 0.0000 0.0000 0.000
f_cyan_ilr_2 11.95 0.00 0.0000 0.0000 0.000
f_JCZ38_qlogis 0.00 16.08 0.0000 0.0000 0.000
log_k1 0.00 0.00 0.9496 0.0000 0.000
log_k2 0.00 0.00 0.0000 0.5846 0.000
g_qlogis 0.00 0.00 0.0000 0.0000 1.719
Starting values for error model parameters:
a.1 b.1
1 1
Results:
Likelihood computed by importance sampling
AIC BIC logLik
2398 2390 -1179
Optimised parameters:
est. lower upper
cyan_0 100.69709 NA NA
log_k_JCZ38 -3.46669 NA NA
log_k_J9Z38 -5.05076 NA NA
log_k_JSE76 -5.55558 NA NA
f_cyan_ilr_1 0.66045 NA NA
f_cyan_ilr_2 0.84275 NA NA
f_JCZ38_qlogis 64.22404 NA NA
log_k1 -2.17715 NA NA
log_k2 -4.55002 NA NA
g_qlogis -0.55920 NA NA
a.1 2.95785 NA NA
b.1 0.04456 NA NA
SD.log_k_JCZ38 1.39881 NA NA
SD.log_k_J9Z38 0.67788 NA NA
SD.log_k_JSE76 0.52603 NA NA
SD.f_cyan_ilr_1 0.32490 NA NA
SD.f_cyan_ilr_2 0.53923 NA NA
SD.f_JCZ38_qlogis 2.75576 NA NA
SD.log_k2 0.30694 NA NA
SD.g_qlogis 0.83619 NA NA
Correlation is not available
Random effects:
est. lower upper
SD.log_k_JCZ38 1.3988 NA NA
SD.log_k_J9Z38 0.6779 NA NA
SD.log_k_JSE76 0.5260 NA NA
SD.f_cyan_ilr_1 0.3249 NA NA
SD.f_cyan_ilr_2 0.5392 NA NA
SD.f_JCZ38_qlogis 2.7558 NA NA
SD.log_k2 0.3069 NA NA
SD.g_qlogis 0.8362 NA NA
Variance model:
est. lower upper
a.1 2.95785 NA NA
b.1 0.04456 NA NA
Backtransformed parameters:
est. lower upper
cyan_0 1.007e+02 NA NA
k_JCZ38 3.122e-02 NA NA
k_J9Z38 6.404e-03 NA NA
k_JSE76 3.866e-03 NA NA
f_cyan_to_JCZ38 6.187e-01 NA NA
f_cyan_to_J9Z38 2.431e-01 NA NA
f_JCZ38_to_JSE76 1.000e+00 NA NA
k1 1.134e-01 NA NA
k2 1.057e-02 NA NA
g 3.637e-01 NA NA
Resulting formation fractions:
ff
cyan_JCZ38 0.6187
cyan_J9Z38 0.2431
cyan_sink 0.1382
JCZ38_JSE76 1.0000
JCZ38_sink 0.0000
Estimated disappearance times:
DT50 DT90 DT50back DT50_k1 DT50_k2
cyan 26.35 175.12 52.72 6.114 65.6
JCZ38 22.20 73.75 NA NA NA
J9Z38 108.23 359.53 NA NA NA
JSE76 179.30 595.62 NA NA NA
saemix version used for fitting: 3.3
mkin version used for pre-fitting: 1.2.10
R version used for fitting: 4.5.0
Date of fit: Mon May 12 20:36:26 2025
Date of summary: Mon May 12 21:06:22 2025
Equations:
d_cyan_free/dt = - k_cyan_free * cyan_free - k_cyan_free_bound *
cyan_free + k_cyan_bound_free * cyan_bound
d_cyan_bound/dt = + k_cyan_free_bound * cyan_free - k_cyan_bound_free *
cyan_bound
d_JCZ38/dt = + f_cyan_free_to_JCZ38 * k_cyan_free * cyan_free - k_JCZ38
* JCZ38
d_J9Z38/dt = + f_cyan_free_to_J9Z38 * k_cyan_free * cyan_free - k_J9Z38
* J9Z38
d_JSE76/dt = + f_JCZ38_to_JSE76 * k_JCZ38 * JCZ38 - k_JSE76 * JSE76
Data:
433 observations of 4 variable(s) grouped in 5 datasets
Model predictions using solution type deSolve
Fitted in 527.639 s
Using 300, 100 iterations and 10 chains
Variance model: Constant variance
Starting values for degradation parameters:
cyan_free_0 log_k_cyan_free log_k_cyan_free_bound
102.0643 -2.8987 -2.7077
log_k_cyan_bound_free log_k_JCZ38 log_k_J9Z38
-3.4717 -3.4008 -5.0024
log_k_JSE76 f_cyan_ilr_1 f_cyan_ilr_2
-5.8613 0.6855 1.2366
f_JCZ38_qlogis
13.7395
Fixed degradation parameter values:
None
Starting values for random effects (square root of initial entries in omega):
cyan_free_0 log_k_cyan_free log_k_cyan_free_bound
cyan_free_0 4.466 0.0000 0.000
log_k_cyan_free 0.000 0.6158 0.000
log_k_cyan_free_bound 0.000 0.0000 1.463
log_k_cyan_bound_free 0.000 0.0000 0.000
log_k_JCZ38 0.000 0.0000 0.000
log_k_J9Z38 0.000 0.0000 0.000
log_k_JSE76 0.000 0.0000 0.000
f_cyan_ilr_1 0.000 0.0000 0.000
f_cyan_ilr_2 0.000 0.0000 0.000
f_JCZ38_qlogis 0.000 0.0000 0.000
log_k_cyan_bound_free log_k_JCZ38 log_k_J9Z38 log_k_JSE76
cyan_free_0 0.000 0.000 0.000 0.000
log_k_cyan_free 0.000 0.000 0.000 0.000
log_k_cyan_free_bound 0.000 0.000 0.000 0.000
log_k_cyan_bound_free 1.058 0.000 0.000 0.000
log_k_JCZ38 0.000 2.382 0.000 0.000
log_k_J9Z38 0.000 0.000 1.595 0.000
log_k_JSE76 0.000 0.000 0.000 1.245
f_cyan_ilr_1 0.000 0.000 0.000 0.000
f_cyan_ilr_2 0.000 0.000 0.000 0.000
f_JCZ38_qlogis 0.000 0.000 0.000 0.000
f_cyan_ilr_1 f_cyan_ilr_2 f_JCZ38_qlogis
cyan_free_0 0.0000 0.00 0.00
log_k_cyan_free 0.0000 0.00 0.00
log_k_cyan_free_bound 0.0000 0.00 0.00
log_k_cyan_bound_free 0.0000 0.00 0.00
log_k_JCZ38 0.0000 0.00 0.00
log_k_J9Z38 0.0000 0.00 0.00
log_k_JSE76 0.0000 0.00 0.00
f_cyan_ilr_1 0.6852 0.00 0.00
f_cyan_ilr_2 0.0000 1.28 0.00
f_JCZ38_qlogis 0.0000 0.00 16.13
Starting values for error model parameters:
a.1
1
Results:
Likelihood computed by importance sampling
AIC BIC logLik
2401 2394 -1181
Optimised parameters:
est. lower upper
cyan_free_0 102.8136 NA NA
log_k_cyan_free -2.7935 NA NA
log_k_cyan_free_bound -2.5440 NA NA
log_k_cyan_bound_free -3.4303 NA NA
log_k_JCZ38 -3.5010 NA NA
log_k_J9Z38 -5.1226 NA NA
log_k_JSE76 -5.6314 NA NA
f_cyan_ilr_1 0.6609 NA NA
f_cyan_ilr_2 0.5085 NA NA
f_JCZ38_qlogis 44.0153 NA NA
a.1 3.2318 NA NA
SD.log_k_cyan_free 0.3211 NA NA
SD.log_k_cyan_free_bound 0.8408 NA NA
SD.log_k_cyan_bound_free 0.5724 NA NA
SD.log_k_JCZ38 1.4925 NA NA
SD.log_k_J9Z38 0.5816 NA NA
SD.log_k_JSE76 0.6037 NA NA
SD.f_cyan_ilr_1 0.3115 NA NA
SD.f_cyan_ilr_2 0.3436 NA NA
SD.f_JCZ38_qlogis 4.8937 NA NA
Correlation is not available
Random effects:
est. lower upper
SD.log_k_cyan_free 0.3211 NA NA
SD.log_k_cyan_free_bound 0.8408 NA NA
SD.log_k_cyan_bound_free 0.5724 NA NA
SD.log_k_JCZ38 1.4925 NA NA
SD.log_k_J9Z38 0.5816 NA NA
SD.log_k_JSE76 0.6037 NA NA
SD.f_cyan_ilr_1 0.3115 NA NA
SD.f_cyan_ilr_2 0.3436 NA NA
SD.f_JCZ38_qlogis 4.8937 NA NA
Variance model:
est. lower upper
a.1 3.232 NA NA
Backtransformed parameters:
est. lower upper
cyan_free_0 1.028e+02 NA NA
k_cyan_free 6.120e-02 NA NA
k_cyan_free_bound 7.855e-02 NA NA
k_cyan_bound_free 3.238e-02 NA NA
k_JCZ38 3.017e-02 NA NA
k_J9Z38 5.961e-03 NA NA
k_JSE76 3.584e-03 NA NA
f_cyan_free_to_JCZ38 5.784e-01 NA NA
f_cyan_free_to_J9Z38 2.271e-01 NA NA
f_JCZ38_to_JSE76 1.000e+00 NA NA
Estimated Eigenvalues of SFORB model(s):
cyan_b1 cyan_b2 cyan_g
0.15973 0.01241 0.33124
Resulting formation fractions:
ff
cyan_free_JCZ38 0.5784
cyan_free_J9Z38 0.2271
cyan_free_sink 0.1945
cyan_free 1.0000
JCZ38_JSE76 1.0000
JCZ38_sink 0.0000
Estimated disappearance times:
DT50 DT90 DT50back DT50_cyan_b1 DT50_cyan_b2
cyan 24.51 153.18 46.11 4.34 55.87
JCZ38 22.98 76.33 NA NA NA
J9Z38 116.28 386.29 NA NA NA
JSE76 193.42 642.53 NA NA NA
saemix version used for fitting: 3.3
mkin version used for pre-fitting: 1.2.10
R version used for fitting: 4.5.0
Date of fit: Mon May 12 20:38:31 2025
Date of summary: Mon May 12 21:06:22 2025
Equations:
d_cyan_free/dt = - k_cyan_free * cyan_free - k_cyan_free_bound *
cyan_free + k_cyan_bound_free * cyan_bound
d_cyan_bound/dt = + k_cyan_free_bound * cyan_free - k_cyan_bound_free *
cyan_bound
d_JCZ38/dt = + f_cyan_free_to_JCZ38 * k_cyan_free * cyan_free - k_JCZ38
* JCZ38
d_J9Z38/dt = + f_cyan_free_to_J9Z38 * k_cyan_free * cyan_free - k_J9Z38
* J9Z38
d_JSE76/dt = + f_JCZ38_to_JSE76 * k_JCZ38 * JCZ38 - k_JSE76 * JSE76
Data:
433 observations of 4 variable(s) grouped in 5 datasets
Model predictions using solution type deSolve
Fitted in 651.67 s
Using 300, 100 iterations and 10 chains
Variance model: Two-component variance function
Starting values for degradation parameters:
cyan_free_0 log_k_cyan_free log_k_cyan_free_bound
101.3964 -2.9881 -2.7949
log_k_cyan_bound_free log_k_JCZ38 log_k_J9Z38
-3.4376 -3.3626 -4.9792
log_k_JSE76 f_cyan_ilr_1 f_cyan_ilr_2
-5.8727 0.6814 6.7399
f_JCZ38_qlogis
13.7395
Fixed degradation parameter values:
None
Starting values for random effects (square root of initial entries in omega):
cyan_free_0 log_k_cyan_free log_k_cyan_free_bound
cyan_free_0 5.317 0.0000 0.000
log_k_cyan_free 0.000 0.7301 0.000
log_k_cyan_free_bound 0.000 0.0000 1.384
log_k_cyan_bound_free 0.000 0.0000 0.000
log_k_JCZ38 0.000 0.0000 0.000
log_k_J9Z38 0.000 0.0000 0.000
log_k_JSE76 0.000 0.0000 0.000
f_cyan_ilr_1 0.000 0.0000 0.000
f_cyan_ilr_2 0.000 0.0000 0.000
f_JCZ38_qlogis 0.000 0.0000 0.000
log_k_cyan_bound_free log_k_JCZ38 log_k_J9Z38 log_k_JSE76
cyan_free_0 0.000 0.000 0.000 0.000
log_k_cyan_free 0.000 0.000 0.000 0.000
log_k_cyan_free_bound 0.000 0.000 0.000 0.000
log_k_cyan_bound_free 1.109 0.000 0.000 0.000
log_k_JCZ38 0.000 2.272 0.000 0.000
log_k_J9Z38 0.000 0.000 1.633 0.000
log_k_JSE76 0.000 0.000 0.000 1.271
f_cyan_ilr_1 0.000 0.000 0.000 0.000
f_cyan_ilr_2 0.000 0.000 0.000 0.000
f_JCZ38_qlogis 0.000 0.000 0.000 0.000
f_cyan_ilr_1 f_cyan_ilr_2 f_JCZ38_qlogis
cyan_free_0 0.0000 0.00 0.00
log_k_cyan_free 0.0000 0.00 0.00
log_k_cyan_free_bound 0.0000 0.00 0.00
log_k_cyan_bound_free 0.0000 0.00 0.00
log_k_JCZ38 0.0000 0.00 0.00
log_k_J9Z38 0.0000 0.00 0.00
log_k_JSE76 0.0000 0.00 0.00
f_cyan_ilr_1 0.6838 0.00 0.00
f_cyan_ilr_2 0.0000 11.69 0.00
f_JCZ38_qlogis 0.0000 0.00 16.13
Starting values for error model parameters:
a.1 b.1
1 1
Results:
Likelihood computed by importance sampling
AIC BIC logLik
2400 2392 -1180
Optimised parameters:
est. lower upper
cyan_free_0 100.56004 NA NA
log_k_cyan_free -3.12657 NA NA
log_k_cyan_free_bound -3.16825 NA NA
log_k_cyan_bound_free -3.66003 NA NA
log_k_JCZ38 -3.47278 NA NA
log_k_J9Z38 -5.06823 NA NA
log_k_JSE76 -5.54327 NA NA
f_cyan_ilr_1 0.66631 NA NA
f_cyan_ilr_2 0.82898 NA NA
f_JCZ38_qlogis 38.31115 NA NA
a.1 2.98352 NA NA
b.1 0.04388 NA NA
SD.log_k_cyan_free 0.49145 NA NA
SD.log_k_cyan_bound_free 0.27347 NA NA
SD.log_k_JCZ38 1.41193 NA NA
SD.log_k_J9Z38 0.66073 NA NA
SD.log_k_JSE76 0.55885 NA NA
SD.f_cyan_ilr_1 0.33020 NA NA
SD.f_cyan_ilr_2 0.51367 NA NA
SD.f_JCZ38_qlogis 5.52122 NA NA
Correlation is not available
Random effects:
est. lower upper
SD.log_k_cyan_free 0.4914 NA NA
SD.log_k_cyan_bound_free 0.2735 NA NA
SD.log_k_JCZ38 1.4119 NA NA
SD.log_k_J9Z38 0.6607 NA NA
SD.log_k_JSE76 0.5589 NA NA
SD.f_cyan_ilr_1 0.3302 NA NA
SD.f_cyan_ilr_2 0.5137 NA NA
SD.f_JCZ38_qlogis 5.5212 NA NA
Variance model:
est. lower upper
a.1 2.98352 NA NA
b.1 0.04388 NA NA
Backtransformed parameters:
est. lower upper
cyan_free_0 1.006e+02 NA NA
k_cyan_free 4.387e-02 NA NA
k_cyan_free_bound 4.208e-02 NA NA
k_cyan_bound_free 2.573e-02 NA NA
k_JCZ38 3.103e-02 NA NA
k_J9Z38 6.294e-03 NA NA
k_JSE76 3.914e-03 NA NA
f_cyan_free_to_JCZ38 6.188e-01 NA NA
f_cyan_free_to_J9Z38 2.412e-01 NA NA
f_JCZ38_to_JSE76 1.000e+00 NA NA
Estimated Eigenvalues of SFORB model(s):
cyan_b1 cyan_b2 cyan_g
0.10044 0.01124 0.36580
Resulting formation fractions:
ff
cyan_free_JCZ38 0.6188
cyan_free_J9Z38 0.2412
cyan_free_sink 0.1400
cyan_free 1.0000
JCZ38_JSE76 1.0000
JCZ38_sink 0.0000
Estimated disappearance times:
DT50 DT90 DT50back DT50_cyan_b1 DT50_cyan_b2
cyan 26.05 164.4 49.48 6.901 61.67
JCZ38 22.34 74.2 NA NA NA
J9Z38 110.14 365.9 NA NA NA
JSE76 177.11 588.3 NA NA NA
saemix version used for fitting: 3.3
mkin version used for pre-fitting: 1.2.10
R version used for fitting: 4.5.0
Date of fit: Mon May 12 20:36:13 2025
Date of summary: Mon May 12 21:06:22 2025
Equations:
d_cyan/dt = - ifelse(time <= tb, k1, k2) * cyan
d_JCZ38/dt = + f_cyan_to_JCZ38 * ifelse(time <= tb, k1, k2) * cyan -
k_JCZ38 * JCZ38
d_J9Z38/dt = + f_cyan_to_J9Z38 * ifelse(time <= tb, k1, k2) * cyan -
k_J9Z38 * J9Z38
d_JSE76/dt = + f_JCZ38_to_JSE76 * k_JCZ38 * JCZ38 - k_JSE76 * JSE76
Data:
433 observations of 4 variable(s) grouped in 5 datasets
Model predictions using solution type deSolve
Fitted in 514.353 s
Using 300, 100 iterations and 10 chains
Variance model: Constant variance
Starting values for degradation parameters:
cyan_0 log_k_JCZ38 log_k_J9Z38 log_k_JSE76 f_cyan_ilr_1
102.8845 -3.4495 -4.9355 -5.6040 0.6468
f_cyan_ilr_2 f_JCZ38_qlogis log_k1 log_k2 log_tb
1.2396 9.7220 -2.9079 -4.1810 1.7813
Fixed degradation parameter values:
None
Starting values for random effects (square root of initial entries in omega):
cyan_0 log_k_JCZ38 log_k_J9Z38 log_k_JSE76 f_cyan_ilr_1
cyan_0 5.406 0.00 0.00 0.000 0.0000
log_k_JCZ38 0.000 2.33 0.00 0.000 0.0000
log_k_J9Z38 0.000 0.00 1.59 0.000 0.0000
log_k_JSE76 0.000 0.00 0.00 1.013 0.0000
f_cyan_ilr_1 0.000 0.00 0.00 0.000 0.6367
f_cyan_ilr_2 0.000 0.00 0.00 0.000 0.0000
f_JCZ38_qlogis 0.000 0.00 0.00 0.000 0.0000
log_k1 0.000 0.00 0.00 0.000 0.0000
log_k2 0.000 0.00 0.00 0.000 0.0000
log_tb 0.000 0.00 0.00 0.000 0.0000
f_cyan_ilr_2 f_JCZ38_qlogis log_k1 log_k2 log_tb
cyan_0 0.000 0.00 0.0000 0.0000 0.0000
log_k_JCZ38 0.000 0.00 0.0000 0.0000 0.0000
log_k_J9Z38 0.000 0.00 0.0000 0.0000 0.0000
log_k_JSE76 0.000 0.00 0.0000 0.0000 0.0000
f_cyan_ilr_1 0.000 0.00 0.0000 0.0000 0.0000
f_cyan_ilr_2 2.038 0.00 0.0000 0.0000 0.0000
f_JCZ38_qlogis 0.000 10.33 0.0000 0.0000 0.0000
log_k1 0.000 0.00 0.7006 0.0000 0.0000
log_k2 0.000 0.00 0.0000 0.8928 0.0000
log_tb 0.000 0.00 0.0000 0.0000 0.6773
Starting values for error model parameters:
a.1
1
Results:
Likelihood computed by importance sampling
AIC BIC logLik
2427 2419 -1194
Optimised parameters:
est. lower upper
cyan_0 101.9660 1.005e+02 1.035e+02
log_k_JCZ38 -3.4698 -4.716e+00 -2.224e+00
log_k_J9Z38 -5.0947 -5.740e+00 -4.450e+00
log_k_JSE76 -5.5977 -6.321e+00 -4.875e+00
f_cyan_ilr_1 0.6595 3.734e-01 9.456e-01
f_cyan_ilr_2 0.5905 1.664e-01 1.015e+00
f_JCZ38_qlogis 25.8627 -4.224e+05 4.225e+05
log_k1 -3.0884 -3.453e+00 -2.723e+00
log_k2 -4.3877 -4.778e+00 -3.998e+00
log_tb 2.3057 1.715e+00 2.896e+00
a.1 3.3228 NA NA
SD.log_k_JCZ38 1.4071 NA NA
SD.log_k_J9Z38 0.5774 NA NA
SD.log_k_JSE76 0.6214 NA NA
SD.f_cyan_ilr_1 0.3058 NA NA
SD.f_cyan_ilr_2 0.3470 NA NA
SD.f_JCZ38_qlogis 0.0644 NA NA
SD.log_k1 0.3994 NA NA
SD.log_k2 0.4373 NA NA
SD.log_tb 0.6419 NA NA
Correlation is not available
Random effects:
est. lower upper
SD.log_k_JCZ38 1.4071 NA NA
SD.log_k_J9Z38 0.5774 NA NA
SD.log_k_JSE76 0.6214 NA NA
SD.f_cyan_ilr_1 0.3058 NA NA
SD.f_cyan_ilr_2 0.3470 NA NA
SD.f_JCZ38_qlogis 0.0644 NA NA
SD.log_k1 0.3994 NA NA
SD.log_k2 0.4373 NA NA
SD.log_tb 0.6419 NA NA
Variance model:
est. lower upper
a.1 3.323 NA NA
Backtransformed parameters:
est. lower upper
cyan_0 1.020e+02 1.005e+02 1.035e+02
k_JCZ38 3.112e-02 8.951e-03 1.082e-01
k_J9Z38 6.129e-03 3.216e-03 1.168e-02
k_JSE76 3.706e-03 1.798e-03 7.639e-03
f_cyan_to_JCZ38 5.890e-01 NA NA
f_cyan_to_J9Z38 2.318e-01 NA NA
f_JCZ38_to_JSE76 1.000e+00 0.000e+00 1.000e+00
k1 4.558e-02 3.164e-02 6.565e-02
k2 1.243e-02 8.417e-03 1.835e-02
tb 1.003e+01 5.557e+00 1.811e+01
Resulting formation fractions:
ff
cyan_JCZ38 5.890e-01
cyan_J9Z38 2.318e-01
cyan_sink 1.793e-01
JCZ38_JSE76 1.000e+00
JCZ38_sink 5.861e-12
Estimated disappearance times:
DT50 DT90 DT50back DT50_k1 DT50_k2
cyan 29.02 158.51 47.72 15.21 55.77
JCZ38 22.27 73.98 NA NA NA
J9Z38 113.09 375.69 NA NA NA
JSE76 187.01 621.23 NA NA NA
Pathway 2
saemix version used for fitting: 3.3
mkin version used for pre-fitting: 1.2.10
R version used for fitting: 4.5.0
Date of fit: Mon May 12 20:47:30 2025
Date of summary: Mon May 12 21:06:22 2025
Equations:
d_cyan/dt = - (alpha/beta) * 1/((time/beta) + 1) * cyan
d_JCZ38/dt = + f_cyan_to_JCZ38 * (alpha/beta) * 1/((time/beta) + 1) *
cyan - k_JCZ38 * JCZ38 + f_JSE76_to_JCZ38 * k_JSE76 * JSE76
d_J9Z38/dt = + f_cyan_to_J9Z38 * (alpha/beta) * 1/((time/beta) + 1) *
cyan - k_J9Z38 * J9Z38
d_JSE76/dt = + f_JCZ38_to_JSE76 * k_JCZ38 * JCZ38 - k_JSE76 * JSE76
Data:
433 observations of 4 variable(s) grouped in 5 datasets
Model predictions using solution type deSolve
Fitted in 505.533 s
Using 300, 100 iterations and 10 chains
Variance model: Two-component variance function
Starting values for degradation parameters:
cyan_0 log_k_JCZ38 log_k_J9Z38 log_k_JSE76 f_cyan_ilr_1
102.4477 -1.8631 -5.1087 -2.5114 0.6826
f_cyan_ilr_2 f_JCZ38_qlogis f_JSE76_qlogis log_alpha log_beta
4.7944 15.9616 13.1566 -0.1564 2.9781
Fixed degradation parameter values:
None
Starting values for random effects (square root of initial entries in omega):
cyan_0 log_k_JCZ38 log_k_J9Z38 log_k_JSE76 f_cyan_ilr_1
cyan_0 7.701 0.000 0.000 0.000 0.0000
log_k_JCZ38 0.000 1.448 0.000 0.000 0.0000
log_k_J9Z38 0.000 0.000 1.724 0.000 0.0000
log_k_JSE76 0.000 0.000 0.000 3.659 0.0000
f_cyan_ilr_1 0.000 0.000 0.000 0.000 0.6356
f_cyan_ilr_2 0.000 0.000 0.000 0.000 0.0000
f_JCZ38_qlogis 0.000 0.000 0.000 0.000 0.0000
f_JSE76_qlogis 0.000 0.000 0.000 0.000 0.0000
log_alpha 0.000 0.000 0.000 0.000 0.0000
log_beta 0.000 0.000 0.000 0.000 0.0000
f_cyan_ilr_2 f_JCZ38_qlogis f_JSE76_qlogis log_alpha log_beta
cyan_0 0.00 0.00 0.00 0.0000 0.0000
log_k_JCZ38 0.00 0.00 0.00 0.0000 0.0000
log_k_J9Z38 0.00 0.00 0.00 0.0000 0.0000
log_k_JSE76 0.00 0.00 0.00 0.0000 0.0000
f_cyan_ilr_1 0.00 0.00 0.00 0.0000 0.0000
f_cyan_ilr_2 10.32 0.00 0.00 0.0000 0.0000
f_JCZ38_qlogis 0.00 12.23 0.00 0.0000 0.0000
f_JSE76_qlogis 0.00 0.00 14.99 0.0000 0.0000
log_alpha 0.00 0.00 0.00 0.3924 0.0000
log_beta 0.00 0.00 0.00 0.0000 0.5639
Starting values for error model parameters:
a.1 b.1
1 1
Results:
Likelihood computed by importance sampling
AIC BIC logLik
2249 2241 -1104
Optimised parameters:
est. lower upper
cyan_0 101.55265 9.920e+01 103.9059
log_k_JCZ38 -2.32302 -2.832e+00 -1.8142
log_k_J9Z38 -5.13082 -5.942e+00 -4.3199
log_k_JSE76 -3.01756 -4.262e+00 -1.7736
f_cyan_ilr_1 0.70850 3.657e-01 1.0513
f_cyan_ilr_2 0.95775 2.612e-01 1.6543
f_JCZ38_qlogis 3.86105 9.248e-01 6.7973
f_JSE76_qlogis 7.51583 -1.120e+02 127.0392
log_alpha -0.15308 -4.508e-01 0.1446
log_beta 2.99165 2.711e+00 3.2720
a.1 2.04034 1.843e+00 2.2382
b.1 0.06924 5.749e-02 0.0810
SD.log_k_JCZ38 0.50818 1.390e-01 0.8774
SD.log_k_J9Z38 0.86597 2.652e-01 1.4667
SD.log_k_JSE76 1.38092 4.864e-01 2.2754
SD.f_cyan_ilr_1 0.38204 1.354e-01 0.6286
SD.f_cyan_ilr_2 0.55129 7.198e-02 1.0306
SD.f_JCZ38_qlogis 1.88457 1.711e-02 3.7520
SD.f_JSE76_qlogis 2.64018 -2.450e+03 2454.9447
SD.log_alpha 0.31860 1.047e-01 0.5325
SD.log_beta 0.24195 1.273e-02 0.4712
Correlation:
cyan_0 l__JCZ3 l__J9Z3 l__JSE7 f_cy__1 f_cy__2 f_JCZ38 f_JSE76
log_k_JCZ38 -0.0235
log_k_J9Z38 -0.0442 0.0047
log_k_JSE76 -0.0023 0.0966 0.0006
f_cyan_ilr_1 -0.0032 0.0070 -0.0536 -0.0001
f_cyan_ilr_2 -0.5189 0.0452 0.1152 0.0013 -0.0304
f_JCZ38_qlogis 0.1088 -0.0848 -0.0240 0.0040 -0.0384 -0.2303
f_JSE76_qlogis -0.0545 0.1315 0.0195 0.0020 0.0252 0.1737 -0.5939
log_alpha -0.0445 0.0056 0.0261 0.0019 -0.0055 0.0586 -0.0239 -0.0284
log_beta -0.2388 0.0163 0.0566 0.0040 -0.0078 0.2183 -0.0714 -0.0332
log_lph
log_k_JCZ38
log_k_J9Z38
log_k_JSE76
f_cyan_ilr_1
f_cyan_ilr_2
f_JCZ38_qlogis
f_JSE76_qlogis
log_alpha
log_beta 0.2135
Random effects:
est. lower upper
SD.log_k_JCZ38 0.5082 1.390e-01 0.8774
SD.log_k_J9Z38 0.8660 2.652e-01 1.4667
SD.log_k_JSE76 1.3809 4.864e-01 2.2754
SD.f_cyan_ilr_1 0.3820 1.354e-01 0.6286
SD.f_cyan_ilr_2 0.5513 7.198e-02 1.0306
SD.f_JCZ38_qlogis 1.8846 1.711e-02 3.7520
SD.f_JSE76_qlogis 2.6402 -2.450e+03 2454.9447
SD.log_alpha 0.3186 1.047e-01 0.5325
SD.log_beta 0.2420 1.273e-02 0.4712
Variance model:
est. lower upper
a.1 2.04034 1.84252 2.238
b.1 0.06924 0.05749 0.081
Backtransformed parameters:
est. lower upper
cyan_0 1.016e+02 9.920e+01 103.9059
k_JCZ38 9.798e-02 5.890e-02 0.1630
k_J9Z38 5.912e-03 2.627e-03 0.0133
k_JSE76 4.892e-02 1.410e-02 0.1697
f_cyan_to_JCZ38 6.432e-01 NA NA
f_cyan_to_J9Z38 2.362e-01 NA NA
f_JCZ38_to_JSE76 9.794e-01 7.160e-01 0.9989
f_JSE76_to_JCZ38 9.995e-01 2.268e-49 1.0000
alpha 8.581e-01 6.371e-01 1.1556
beta 1.992e+01 1.505e+01 26.3646
Resulting formation fractions:
ff
cyan_JCZ38 0.6432301
cyan_J9Z38 0.2361657
cyan_sink 0.1206042
JCZ38_JSE76 0.9793879
JCZ38_sink 0.0206121
JSE76_JCZ38 0.9994559
JSE76_sink 0.0005441
Estimated disappearance times:
DT50 DT90 DT50back
cyan 24.759 271.61 81.76
JCZ38 7.075 23.50 NA
J9Z38 117.249 389.49 NA
JSE76 14.169 47.07 NA
saemix version used for fitting: 3.3
mkin version used for pre-fitting: 1.2.10
R version used for fitting: 4.5.0
Date of fit: Mon May 12 20:48:20 2025
Date of summary: Mon May 12 21:06:22 2025
Equations:
d_cyan/dt = - ((k1 * g * exp(-k1 * time) + k2 * (1 - g) * exp(-k2 *
time)) / (g * exp(-k1 * time) + (1 - g) * exp(-k2 * time)))
* cyan
d_JCZ38/dt = + f_cyan_to_JCZ38 * ((k1 * g * exp(-k1 * time) + k2 * (1 -
g) * exp(-k2 * time)) / (g * exp(-k1 * time) + (1 - g) *
exp(-k2 * time))) * cyan - k_JCZ38 * JCZ38 +
f_JSE76_to_JCZ38 * k_JSE76 * JSE76
d_J9Z38/dt = + f_cyan_to_J9Z38 * ((k1 * g * exp(-k1 * time) + k2 * (1 -
g) * exp(-k2 * time)) / (g * exp(-k1 * time) + (1 - g) *
exp(-k2 * time))) * cyan - k_J9Z38 * J9Z38
d_JSE76/dt = + f_JCZ38_to_JSE76 * k_JCZ38 * JCZ38 - k_JSE76 * JSE76
Data:
433 observations of 4 variable(s) grouped in 5 datasets
Model predictions using solution type deSolve
Fitted in 555.413 s
Using 300, 100 iterations and 10 chains
Variance model: Constant variance
Starting values for degradation parameters:
cyan_0 log_k_JCZ38 log_k_J9Z38 log_k_JSE76 f_cyan_ilr_1
102.4380 -2.3107 -5.3123 -3.7120 0.6757
f_cyan_ilr_2 f_JCZ38_qlogis f_JSE76_qlogis log_k1 log_k2
1.1439 13.1194 12.3492 -1.9317 -4.4557
g_qlogis
-0.5644
Fixed degradation parameter values:
None
Starting values for random effects (square root of initial entries in omega):
cyan_0 log_k_JCZ38 log_k_J9Z38 log_k_JSE76 f_cyan_ilr_1
cyan_0 4.591 0.0000 0.000 0.0 0.0000
log_k_JCZ38 0.000 0.7966 0.000 0.0 0.0000
log_k_J9Z38 0.000 0.0000 1.561 0.0 0.0000
log_k_JSE76 0.000 0.0000 0.000 0.8 0.0000
f_cyan_ilr_1 0.000 0.0000 0.000 0.0 0.6349
f_cyan_ilr_2 0.000 0.0000 0.000 0.0 0.0000
f_JCZ38_qlogis 0.000 0.0000 0.000 0.0 0.0000
f_JSE76_qlogis 0.000 0.0000 0.000 0.0 0.0000
log_k1 0.000 0.0000 0.000 0.0 0.0000
log_k2 0.000 0.0000 0.000 0.0 0.0000
g_qlogis 0.000 0.0000 0.000 0.0 0.0000
f_cyan_ilr_2 f_JCZ38_qlogis f_JSE76_qlogis log_k1 log_k2
cyan_0 0.000 0.00 0.00 0.000 0.0000
log_k_JCZ38 0.000 0.00 0.00 0.000 0.0000
log_k_J9Z38 0.000 0.00 0.00 0.000 0.0000
log_k_JSE76 0.000 0.00 0.00 0.000 0.0000
f_cyan_ilr_1 0.000 0.00 0.00 0.000 0.0000
f_cyan_ilr_2 1.797 0.00 0.00 0.000 0.0000
f_JCZ38_qlogis 0.000 13.86 0.00 0.000 0.0000
f_JSE76_qlogis 0.000 0.00 13.91 0.000 0.0000
log_k1 0.000 0.00 0.00 1.106 0.0000
log_k2 0.000 0.00 0.00 0.000 0.6141
g_qlogis 0.000 0.00 0.00 0.000 0.0000
g_qlogis
cyan_0 0.000
log_k_JCZ38 0.000
log_k_J9Z38 0.000
log_k_JSE76 0.000
f_cyan_ilr_1 0.000
f_cyan_ilr_2 0.000
f_JCZ38_qlogis 0.000
f_JSE76_qlogis 0.000
log_k1 0.000
log_k2 0.000
g_qlogis 1.595
Starting values for error model parameters:
a.1
1
Results:
Likelihood computed by importance sampling
AIC BIC logLik
2288 2280 -1122
Optimised parameters:
est. lower upper
cyan_0 102.7204 1.014e+02 1.040e+02
log_k_JCZ38 -2.8925 -4.044e+00 -1.741e+00
log_k_J9Z38 -5.1430 -5.828e+00 -4.457e+00
log_k_JSE76 -3.5577 -4.174e+00 -2.941e+00
f_cyan_ilr_1 0.6929 3.788e-01 1.007e+00
f_cyan_ilr_2 0.6066 5.342e-02 1.160e+00
f_JCZ38_qlogis 9.8071 -2.819e+03 2.838e+03
f_JSE76_qlogis 2.2229 5.684e-01 3.877e+00
log_k1 -1.9339 -2.609e+00 -1.258e+00
log_k2 -4.4709 -4.935e+00 -4.007e+00
g_qlogis -0.4987 -1.373e+00 3.757e-01
a.1 2.7368 2.545e+00 2.928e+00
SD.log_k_JCZ38 1.2747 4.577e-01 2.092e+00
SD.log_k_J9Z38 0.6758 1.418e-01 1.210e+00
SD.log_k_JSE76 0.5869 1.169e-01 1.057e+00
SD.f_cyan_ilr_1 0.3392 1.161e-01 5.622e-01
SD.f_cyan_ilr_2 0.4200 8.501e-02 7.550e-01
SD.f_JCZ38_qlogis 0.8511 -1.137e+06 1.137e+06
SD.f_JSE76_qlogis 0.3767 -5.238e-01 1.277e+00
SD.log_k1 0.7475 2.601e-01 1.235e+00
SD.log_k2 0.5179 1.837e-01 8.521e-01
SD.g_qlogis 0.9817 3.553e-01 1.608e+00
Correlation:
cyan_0 l__JCZ3 l__J9Z3 l__JSE7 f_cy__1 f_cy__2 f_JCZ38 f_JSE76
log_k_JCZ38 -0.0351
log_k_J9Z38 -0.0541 0.0043
log_k_JSE76 -0.0078 0.0900 -0.0014
f_cyan_ilr_1 -0.0249 0.0268 -0.0962 0.0000
f_cyan_ilr_2 -0.3560 0.0848 0.1545 -0.0022 0.0463
f_JCZ38_qlogis 0.2005 -0.1226 -0.0347 0.0514 -0.1840 -0.5906
f_JSE76_qlogis -0.1638 0.1307 0.0266 0.0001 0.1645 0.5181 -0.9297
log_k1 0.0881 -0.0071 0.0005 -0.0070 -0.0064 -0.0346 0.0316 -0.0341
log_k2 0.0238 -0.0003 0.0082 -0.0022 -0.0017 -0.0017 -0.0002 -0.0076
g_qlogis 0.0198 -0.0002 -0.0109 0.0034 0.0017 -0.0176 0.0044 0.0051
log_k1 log_k2
log_k_JCZ38
log_k_J9Z38
log_k_JSE76
f_cyan_ilr_1
f_cyan_ilr_2
f_JCZ38_qlogis
f_JSE76_qlogis
log_k1
log_k2 0.0276
g_qlogis -0.0283 -0.0309
Random effects:
est. lower upper
SD.log_k_JCZ38 1.2747 4.577e-01 2.092e+00
SD.log_k_J9Z38 0.6758 1.418e-01 1.210e+00
SD.log_k_JSE76 0.5869 1.169e-01 1.057e+00
SD.f_cyan_ilr_1 0.3392 1.161e-01 5.622e-01
SD.f_cyan_ilr_2 0.4200 8.501e-02 7.550e-01
SD.f_JCZ38_qlogis 0.8511 -1.137e+06 1.137e+06
SD.f_JSE76_qlogis 0.3767 -5.238e-01 1.277e+00
SD.log_k1 0.7475 2.601e-01 1.235e+00
SD.log_k2 0.5179 1.837e-01 8.521e-01
SD.g_qlogis 0.9817 3.553e-01 1.608e+00
Variance model:
est. lower upper
a.1 2.737 2.545 2.928
Backtransformed parameters:
est. lower upper
cyan_0 102.72037 1.014e+02 104.00464
k_JCZ38 0.05544 1.752e-02 0.17539
k_J9Z38 0.00584 2.942e-03 0.01159
k_JSE76 0.02850 1.539e-02 0.05279
f_cyan_to_JCZ38 0.59995 NA NA
f_cyan_to_J9Z38 0.22519 NA NA
f_JCZ38_to_JSE76 0.99994 0.000e+00 1.00000
f_JSE76_to_JCZ38 0.90229 6.384e-01 0.97971
k1 0.14459 7.357e-02 0.28414
k2 0.01144 7.192e-03 0.01819
g 0.37784 2.021e-01 0.59284
Resulting formation fractions:
ff
cyan_JCZ38 5.999e-01
cyan_J9Z38 2.252e-01
cyan_sink 1.749e-01
JCZ38_JSE76 9.999e-01
JCZ38_sink 5.506e-05
JSE76_JCZ38 9.023e-01
JSE76_sink 9.771e-02
Estimated disappearance times:
DT50 DT90 DT50back DT50_k1 DT50_k2
cyan 21.93 159.83 48.11 4.794 60.6
JCZ38 12.50 41.53 NA NA NA
J9Z38 118.69 394.27 NA NA NA
JSE76 24.32 80.78 NA NA NA
saemix version used for fitting: 3.3
mkin version used for pre-fitting: 1.2.10
R version used for fitting: 4.5.0
Date of fit: Mon May 12 20:51:02 2025
Date of summary: Mon May 12 21:06:22 2025
Equations:
d_cyan/dt = - ((k1 * g * exp(-k1 * time) + k2 * (1 - g) * exp(-k2 *
time)) / (g * exp(-k1 * time) + (1 - g) * exp(-k2 * time)))
* cyan
d_JCZ38/dt = + f_cyan_to_JCZ38 * ((k1 * g * exp(-k1 * time) + k2 * (1 -
g) * exp(-k2 * time)) / (g * exp(-k1 * time) + (1 - g) *
exp(-k2 * time))) * cyan - k_JCZ38 * JCZ38 +
f_JSE76_to_JCZ38 * k_JSE76 * JSE76
d_J9Z38/dt = + f_cyan_to_J9Z38 * ((k1 * g * exp(-k1 * time) + k2 * (1 -
g) * exp(-k2 * time)) / (g * exp(-k1 * time) + (1 - g) *
exp(-k2 * time))) * cyan - k_J9Z38 * J9Z38
d_JSE76/dt = + f_JCZ38_to_JSE76 * k_JCZ38 * JCZ38 - k_JSE76 * JSE76
Data:
433 observations of 4 variable(s) grouped in 5 datasets
Model predictions using solution type deSolve
Fitted in 717.675 s
Using 300, 100 iterations and 10 chains
Variance model: Two-component variance function
Starting values for degradation parameters:
cyan_0 log_k_JCZ38 log_k_J9Z38 log_k_JSE76 f_cyan_ilr_1
101.7393 -1.4493 -5.0118 -2.1269 0.6720
f_cyan_ilr_2 f_JCZ38_qlogis f_JSE76_qlogis log_k1 log_k2
7.3362 13.4423 13.2659 -2.0061 -4.5527
g_qlogis
-0.5806
Fixed degradation parameter values:
None
Starting values for random effects (square root of initial entries in omega):
cyan_0 log_k_JCZ38 log_k_J9Z38 log_k_JSE76 f_cyan_ilr_1
cyan_0 5.604 0.00 0.000 0.000 0.0000
log_k_JCZ38 0.000 2.77 0.000 0.000 0.0000
log_k_J9Z38 0.000 0.00 1.662 0.000 0.0000
log_k_JSE76 0.000 0.00 0.000 5.021 0.0000
f_cyan_ilr_1 0.000 0.00 0.000 0.000 0.6519
f_cyan_ilr_2 0.000 0.00 0.000 0.000 0.0000
f_JCZ38_qlogis 0.000 0.00 0.000 0.000 0.0000
f_JSE76_qlogis 0.000 0.00 0.000 0.000 0.0000
log_k1 0.000 0.00 0.000 0.000 0.0000
log_k2 0.000 0.00 0.000 0.000 0.0000
g_qlogis 0.000 0.00 0.000 0.000 0.0000
f_cyan_ilr_2 f_JCZ38_qlogis f_JSE76_qlogis log_k1 log_k2
cyan_0 0.00 0.00 0.00 0.0000 0.0000
log_k_JCZ38 0.00 0.00 0.00 0.0000 0.0000
log_k_J9Z38 0.00 0.00 0.00 0.0000 0.0000
log_k_JSE76 0.00 0.00 0.00 0.0000 0.0000
f_cyan_ilr_1 0.00 0.00 0.00 0.0000 0.0000
f_cyan_ilr_2 13.37 0.00 0.00 0.0000 0.0000
f_JCZ38_qlogis 0.00 14.21 0.00 0.0000 0.0000
f_JSE76_qlogis 0.00 0.00 14.58 0.0000 0.0000
log_k1 0.00 0.00 0.00 0.8453 0.0000
log_k2 0.00 0.00 0.00 0.0000 0.5969
g_qlogis 0.00 0.00 0.00 0.0000 0.0000
g_qlogis
cyan_0 0.00
log_k_JCZ38 0.00
log_k_J9Z38 0.00
log_k_JSE76 0.00
f_cyan_ilr_1 0.00
f_cyan_ilr_2 0.00
f_JCZ38_qlogis 0.00
f_JSE76_qlogis 0.00
log_k1 0.00
log_k2 0.00
g_qlogis 1.69
Starting values for error model parameters:
a.1 b.1
1 1
Results:
Likelihood computed by importance sampling
AIC BIC logLik
2234 2226 -1095
Optimised parameters:
est. lower upper
cyan_0 101.25496 99.14662 103.36331
log_k_JCZ38 -2.55593 -3.32972 -1.78215
log_k_J9Z38 -5.07103 -5.85423 -4.28783
log_k_JSE76 -3.25468 -4.17577 -2.33360
f_cyan_ilr_1 0.70139 0.35924 1.04355
f_cyan_ilr_2 1.07712 0.17789 1.97636
f_JCZ38_qlogis 3.57483 0.05990 7.08976
f_JSE76_qlogis 4.54884 -7.25628 16.35395
log_k1 -2.38201 -2.51639 -2.24763
log_k2 -4.66741 -4.91865 -4.41617
g_qlogis -0.28446 -1.14192 0.57300
a.1 2.05925 1.86481 2.25369
b.1 0.06172 0.05062 0.07282
SD.log_k_JCZ38 0.81137 0.25296 1.36977
SD.log_k_J9Z38 0.83542 0.25395 1.41689
SD.log_k_JSE76 0.97903 0.30100 1.65707
SD.f_cyan_ilr_1 0.37878 0.13374 0.62382
SD.f_cyan_ilr_2 0.67274 0.10102 1.24446
SD.f_JCZ38_qlogis 1.35327 -0.42359 3.13012
SD.f_JSE76_qlogis 1.43956 -19.14972 22.02884
SD.log_k2 0.25329 0.07521 0.43138
SD.g_qlogis 0.95167 0.35149 1.55184
Correlation:
cyan_0 l__JCZ3 l__J9Z3 l__JSE7 f_cy__1 f_cy__2 f_JCZ38 f_JSE76
log_k_JCZ38 -0.0265
log_k_J9Z38 -0.0392 0.0024
log_k_JSE76 0.0011 0.1220 -0.0016
f_cyan_ilr_1 -0.0161 0.0217 -0.0552 0.0034
f_cyan_ilr_2 -0.4718 0.0829 0.1102 0.0042 0.0095
f_JCZ38_qlogis 0.1609 -0.1318 -0.0277 0.0081 -0.1040 -0.4559
f_JSE76_qlogis -0.1289 0.1494 0.0219 0.0012 0.1004 0.4309 -0.8543
log_k1 0.2618 -0.0739 -0.0167 -0.0148 -0.0444 -0.2768 0.3518 -0.3818
log_k2 0.0603 -0.0217 0.0174 -0.0058 -0.0197 -0.0533 0.0923 -0.1281
g_qlogis 0.0362 0.0115 -0.0111 0.0040 0.0095 -0.0116 -0.0439 0.0651
log_k1 log_k2
log_k_JCZ38
log_k_J9Z38
log_k_JSE76
f_cyan_ilr_1
f_cyan_ilr_2
f_JCZ38_qlogis
f_JSE76_qlogis
log_k1
log_k2 0.3269
g_qlogis -0.1656 -0.0928
Random effects:
est. lower upper
SD.log_k_JCZ38 0.8114 0.25296 1.3698
SD.log_k_J9Z38 0.8354 0.25395 1.4169
SD.log_k_JSE76 0.9790 0.30100 1.6571
SD.f_cyan_ilr_1 0.3788 0.13374 0.6238
SD.f_cyan_ilr_2 0.6727 0.10102 1.2445
SD.f_JCZ38_qlogis 1.3533 -0.42359 3.1301
SD.f_JSE76_qlogis 1.4396 -19.14972 22.0288
SD.log_k2 0.2533 0.07521 0.4314
SD.g_qlogis 0.9517 0.35149 1.5518
Variance model:
est. lower upper
a.1 2.05925 1.86481 2.25369
b.1 0.06172 0.05062 0.07282
Backtransformed parameters:
est. lower upper
cyan_0 1.013e+02 9.915e+01 103.36331
k_JCZ38 7.762e-02 3.580e-02 0.16828
k_J9Z38 6.276e-03 2.868e-03 0.01373
k_JSE76 3.859e-02 1.536e-02 0.09695
f_cyan_to_JCZ38 6.520e-01 NA NA
f_cyan_to_J9Z38 2.418e-01 NA NA
f_JCZ38_to_JSE76 9.727e-01 5.150e-01 0.99917
f_JSE76_to_JCZ38 9.895e-01 7.052e-04 1.00000
k1 9.236e-02 8.075e-02 0.10565
k2 9.397e-03 7.309e-03 0.01208
g 4.294e-01 2.420e-01 0.63945
Resulting formation fractions:
ff
cyan_JCZ38 0.65203
cyan_J9Z38 0.24181
cyan_sink 0.10616
JCZ38_JSE76 0.97274
JCZ38_sink 0.02726
JSE76_JCZ38 0.98953
JSE76_sink 0.01047
Estimated disappearance times:
DT50 DT90 DT50back DT50_k1 DT50_k2
cyan 24.26 185.34 55.79 7.504 73.77
JCZ38 8.93 29.66 NA NA NA
J9Z38 110.45 366.89 NA NA NA
JSE76 17.96 59.66 NA NA NA
saemix version used for fitting: 3.3
mkin version used for pre-fitting: 1.2.10
R version used for fitting: 4.5.0
Date of fit: Mon May 12 20:48:15 2025
Date of summary: Mon May 12 21:06:22 2025
Equations:
d_cyan_free/dt = - k_cyan_free * cyan_free - k_cyan_free_bound *
cyan_free + k_cyan_bound_free * cyan_bound
d_cyan_bound/dt = + k_cyan_free_bound * cyan_free - k_cyan_bound_free *
cyan_bound
d_JCZ38/dt = + f_cyan_free_to_JCZ38 * k_cyan_free * cyan_free - k_JCZ38
* JCZ38 + f_JSE76_to_JCZ38 * k_JSE76 * JSE76
d_J9Z38/dt = + f_cyan_free_to_J9Z38 * k_cyan_free * cyan_free - k_J9Z38
* J9Z38
d_JSE76/dt = + f_JCZ38_to_JSE76 * k_JCZ38 * JCZ38 - k_JSE76 * JSE76
Data:
433 observations of 4 variable(s) grouped in 5 datasets
Model predictions using solution type deSolve
Fitted in 550.772 s
Using 300, 100 iterations and 10 chains
Variance model: Constant variance
Starting values for degradation parameters:
cyan_free_0 log_k_cyan_free log_k_cyan_free_bound
102.4395 -2.7673 -2.8942
log_k_cyan_bound_free log_k_JCZ38 log_k_J9Z38
-3.6201 -2.3107 -5.3123
log_k_JSE76 f_cyan_ilr_1 f_cyan_ilr_2
-3.7120 0.6754 1.1448
f_JCZ38_qlogis f_JSE76_qlogis
14.8408 15.4734
Fixed degradation parameter values:
None
Starting values for random effects (square root of initial entries in omega):
cyan_free_0 log_k_cyan_free log_k_cyan_free_bound
cyan_free_0 4.589 0.0000 0.00
log_k_cyan_free 0.000 0.4849 0.00
log_k_cyan_free_bound 0.000 0.0000 1.62
log_k_cyan_bound_free 0.000 0.0000 0.00
log_k_JCZ38 0.000 0.0000 0.00
log_k_J9Z38 0.000 0.0000 0.00
log_k_JSE76 0.000 0.0000 0.00
f_cyan_ilr_1 0.000 0.0000 0.00
f_cyan_ilr_2 0.000 0.0000 0.00
f_JCZ38_qlogis 0.000 0.0000 0.00
f_JSE76_qlogis 0.000 0.0000 0.00
log_k_cyan_bound_free log_k_JCZ38 log_k_J9Z38 log_k_JSE76
cyan_free_0 0.000 0.0000 0.000 0.0
log_k_cyan_free 0.000 0.0000 0.000 0.0
log_k_cyan_free_bound 0.000 0.0000 0.000 0.0
log_k_cyan_bound_free 1.197 0.0000 0.000 0.0
log_k_JCZ38 0.000 0.7966 0.000 0.0
log_k_J9Z38 0.000 0.0000 1.561 0.0
log_k_JSE76 0.000 0.0000 0.000 0.8
f_cyan_ilr_1 0.000 0.0000 0.000 0.0
f_cyan_ilr_2 0.000 0.0000 0.000 0.0
f_JCZ38_qlogis 0.000 0.0000 0.000 0.0
f_JSE76_qlogis 0.000 0.0000 0.000 0.0
f_cyan_ilr_1 f_cyan_ilr_2 f_JCZ38_qlogis f_JSE76_qlogis
cyan_free_0 0.0000 0.000 0.0 0.00
log_k_cyan_free 0.0000 0.000 0.0 0.00
log_k_cyan_free_bound 0.0000 0.000 0.0 0.00
log_k_cyan_bound_free 0.0000 0.000 0.0 0.00
log_k_JCZ38 0.0000 0.000 0.0 0.00
log_k_J9Z38 0.0000 0.000 0.0 0.00
log_k_JSE76 0.0000 0.000 0.0 0.00
f_cyan_ilr_1 0.6349 0.000 0.0 0.00
f_cyan_ilr_2 0.0000 1.797 0.0 0.00
f_JCZ38_qlogis 0.0000 0.000 15.6 0.00
f_JSE76_qlogis 0.0000 0.000 0.0 17.52
Starting values for error model parameters:
a.1
1
Results:
Likelihood computed by importance sampling
AIC BIC logLik
2283 2275 -1120
Optimised parameters:
est. lower upper
cyan_free_0 102.6517 101.40815 103.8952
log_k_cyan_free -2.8729 -3.18649 -2.5593
log_k_cyan_free_bound -2.7803 -3.60525 -1.9552
log_k_cyan_bound_free -3.5845 -4.16644 -3.0026
log_k_JCZ38 -2.3411 -2.89698 -1.7852
log_k_J9Z38 -5.2487 -6.01271 -4.4847
log_k_JSE76 -3.0259 -4.28274 -1.7690
f_cyan_ilr_1 0.7289 0.38214 1.0756
f_cyan_ilr_2 0.6891 0.18277 1.1954
f_JCZ38_qlogis 4.2162 0.47015 7.9622
f_JSE76_qlogis 5.8911 -20.19088 31.9730
a.1 2.7159 2.52587 2.9060
SD.log_k_cyan_free 0.3354 0.10979 0.5610
SD.log_k_cyan_free_bound 0.9061 0.30969 1.5025
SD.log_k_cyan_bound_free 0.6376 0.21229 1.0628
SD.log_k_JCZ38 0.5499 0.14533 0.9545
SD.log_k_J9Z38 0.7457 0.15106 1.3404
SD.log_k_JSE76 1.3822 0.47329 2.2912
SD.f_cyan_ilr_1 0.3820 0.13280 0.6313
SD.f_cyan_ilr_2 0.4317 0.06803 0.7953
SD.f_JCZ38_qlogis 1.8258 -0.25423 3.9059
SD.f_JSE76_qlogis 2.2348 -83.33679 87.8065
Correlation:
cyn_f_0 lg_k_c_ lg_k_cyn_f_ lg_k_cyn_b_ l__JCZ3 l__J9Z3
log_k_cyan_free 0.1944
log_k_cyan_free_bound 0.0815 0.0814
log_k_cyan_bound_free 0.0106 0.0426 0.0585
log_k_JCZ38 -0.0231 -0.0106 -0.0089 -0.0051
log_k_J9Z38 -0.0457 -0.0108 0.0019 0.0129 0.0032
log_k_JSE76 -0.0054 -0.0024 -0.0017 -0.0005 0.1108 0.0009
f_cyan_ilr_1 0.0051 -0.0005 -0.0035 -0.0056 0.0131 -0.0967
f_cyan_ilr_2 -0.3182 -0.0771 -0.0309 -0.0038 0.0680 0.1643
f_JCZ38_qlogis 0.0834 0.0369 0.0302 0.0172 -0.1145 -0.0204
f_JSE76_qlogis -0.0553 -0.0365 -0.0441 -0.0414 0.1579 0.0175
l__JSE7 f_cy__1 f_cy__2 f_JCZ38
log_k_cyan_free
log_k_cyan_free_bound
log_k_cyan_bound_free
log_k_JCZ38
log_k_J9Z38
log_k_JSE76
f_cyan_ilr_1 -0.0002
f_cyan_ilr_2 0.0020 -0.0415
f_JCZ38_qlogis 0.0052 -0.0665 -0.3437
f_JSE76_qlogis 0.0066 0.0635 0.3491 -0.7487
Random effects:
est. lower upper
SD.log_k_cyan_free 0.3354 0.10979 0.5610
SD.log_k_cyan_free_bound 0.9061 0.30969 1.5025
SD.log_k_cyan_bound_free 0.6376 0.21229 1.0628
SD.log_k_JCZ38 0.5499 0.14533 0.9545
SD.log_k_J9Z38 0.7457 0.15106 1.3404
SD.log_k_JSE76 1.3822 0.47329 2.2912
SD.f_cyan_ilr_1 0.3820 0.13280 0.6313
SD.f_cyan_ilr_2 0.4317 0.06803 0.7953
SD.f_JCZ38_qlogis 1.8258 -0.25423 3.9059
SD.f_JSE76_qlogis 2.2348 -83.33679 87.8065
Variance model:
est. lower upper
a.1 2.716 2.526 2.906
Backtransformed parameters:
est. lower upper
cyan_free_0 1.027e+02 1.014e+02 103.89517
k_cyan_free 5.654e-02 4.132e-02 0.07736
k_cyan_free_bound 6.202e-02 2.718e-02 0.14153
k_cyan_bound_free 2.775e-02 1.551e-02 0.04966
k_JCZ38 9.622e-02 5.519e-02 0.16777
k_J9Z38 5.254e-03 2.447e-03 0.01128
k_JSE76 4.852e-02 1.380e-02 0.17051
f_cyan_free_to_JCZ38 6.197e-01 5.643e-01 0.84429
f_cyan_free_to_J9Z38 2.211e-01 5.643e-01 0.84429
f_JCZ38_to_JSE76 9.855e-01 6.154e-01 0.99965
f_JSE76_to_JCZ38 9.972e-01 1.703e-09 1.00000
Estimated Eigenvalues of SFORB model(s):
cyan_b1 cyan_b2 cyan_g
0.13466 0.01165 0.36490
Resulting formation fractions:
ff
cyan_free_JCZ38 0.619745
cyan_free_J9Z38 0.221083
cyan_free_sink 0.159172
cyan_free 1.000000
JCZ38_JSE76 0.985460
JCZ38_sink 0.014540
JSE76_JCZ38 0.997244
JSE76_sink 0.002756
Estimated disappearance times:
DT50 DT90 DT50back DT50_cyan_b1 DT50_cyan_b2
cyan 23.293 158.67 47.77 5.147 59.5
JCZ38 7.203 23.93 NA NA NA
J9Z38 131.918 438.22 NA NA NA
JSE76 14.287 47.46 NA NA NA
saemix version used for fitting: 3.3
mkin version used for pre-fitting: 1.2.10
R version used for fitting: 4.5.0
Date of fit: Mon May 12 20:50:59 2025
Date of summary: Mon May 12 21:06:22 2025
Equations:
d_cyan_free/dt = - k_cyan_free * cyan_free - k_cyan_free_bound *
cyan_free + k_cyan_bound_free * cyan_bound
d_cyan_bound/dt = + k_cyan_free_bound * cyan_free - k_cyan_bound_free *
cyan_bound
d_JCZ38/dt = + f_cyan_free_to_JCZ38 * k_cyan_free * cyan_free - k_JCZ38
* JCZ38 + f_JSE76_to_JCZ38 * k_JSE76 * JSE76
d_J9Z38/dt = + f_cyan_free_to_J9Z38 * k_cyan_free * cyan_free - k_J9Z38
* J9Z38
d_JSE76/dt = + f_JCZ38_to_JSE76 * k_JCZ38 * JCZ38 - k_JSE76 * JSE76
Data:
433 observations of 4 variable(s) grouped in 5 datasets
Model predictions using solution type deSolve
Fitted in 714.48 s
Using 300, 100 iterations and 10 chains
Variance model: Two-component variance function
Starting values for degradation parameters:
cyan_free_0 log_k_cyan_free log_k_cyan_free_bound
101.7511 -2.8370 -3.0162
log_k_cyan_bound_free log_k_JCZ38 log_k_J9Z38
-3.6600 -2.2988 -5.3129
log_k_JSE76 f_cyan_ilr_1 f_cyan_ilr_2
-3.6991 0.6722 4.8596
f_JCZ38_qlogis f_JSE76_qlogis
13.4678 14.2149
Fixed degradation parameter values:
None
Starting values for random effects (square root of initial entries in omega):
cyan_free_0 log_k_cyan_free log_k_cyan_free_bound
cyan_free_0 5.629 0.000 0.000
log_k_cyan_free 0.000 0.446 0.000
log_k_cyan_free_bound 0.000 0.000 1.449
log_k_cyan_bound_free 0.000 0.000 0.000
log_k_JCZ38 0.000 0.000 0.000
log_k_J9Z38 0.000 0.000 0.000
log_k_JSE76 0.000 0.000 0.000
f_cyan_ilr_1 0.000 0.000 0.000
f_cyan_ilr_2 0.000 0.000 0.000
f_JCZ38_qlogis 0.000 0.000 0.000
f_JSE76_qlogis 0.000 0.000 0.000
log_k_cyan_bound_free log_k_JCZ38 log_k_J9Z38 log_k_JSE76
cyan_free_0 0.000 0.0000 0.000 0.0000
log_k_cyan_free 0.000 0.0000 0.000 0.0000
log_k_cyan_free_bound 0.000 0.0000 0.000 0.0000
log_k_cyan_bound_free 1.213 0.0000 0.000 0.0000
log_k_JCZ38 0.000 0.7801 0.000 0.0000
log_k_J9Z38 0.000 0.0000 1.575 0.0000
log_k_JSE76 0.000 0.0000 0.000 0.8078
f_cyan_ilr_1 0.000 0.0000 0.000 0.0000
f_cyan_ilr_2 0.000 0.0000 0.000 0.0000
f_JCZ38_qlogis 0.000 0.0000 0.000 0.0000
f_JSE76_qlogis 0.000 0.0000 0.000 0.0000
f_cyan_ilr_1 f_cyan_ilr_2 f_JCZ38_qlogis f_JSE76_qlogis
cyan_free_0 0.0000 0.000 0.00 0.00
log_k_cyan_free 0.0000 0.000 0.00 0.00
log_k_cyan_free_bound 0.0000 0.000 0.00 0.00
log_k_cyan_bound_free 0.0000 0.000 0.00 0.00
log_k_JCZ38 0.0000 0.000 0.00 0.00
log_k_J9Z38 0.0000 0.000 0.00 0.00
log_k_JSE76 0.0000 0.000 0.00 0.00
f_cyan_ilr_1 0.6518 0.000 0.00 0.00
f_cyan_ilr_2 0.0000 9.981 0.00 0.00
f_JCZ38_qlogis 0.0000 0.000 14.26 0.00
f_JSE76_qlogis 0.0000 0.000 0.00 16.17
Starting values for error model parameters:
a.1 b.1
1 1
Results:
Likelihood computed by importance sampling
AIC BIC logLik
2240 2231 -1098
Optimised parameters:
est. lower upper
cyan_free_0 100.73014 9.873e+01 1.027e+02
log_k_cyan_free -3.19634 -3.641e+00 -2.752e+00
log_k_cyan_free_bound -3.43533 -3.674e+00 -3.197e+00
log_k_cyan_bound_free -3.83282 -4.163e+00 -3.503e+00
log_k_JCZ38 -2.51065 -3.225e+00 -1.796e+00
log_k_J9Z38 -5.02539 -5.825e+00 -4.226e+00
log_k_JSE76 -3.24777 -4.163e+00 -2.333e+00
f_cyan_ilr_1 0.70640 3.562e-01 1.057e+00
f_cyan_ilr_2 1.42704 3.170e-01 2.537e+00
f_JCZ38_qlogis 2.84779 1.042e+00 4.654e+00
f_JSE76_qlogis 8.63674 -6.407e+02 6.580e+02
a.1 2.07082 1.877e+00 2.265e+00
b.1 0.06227 5.098e-02 7.355e-02
SD.log_k_cyan_free 0.49674 1.865e-01 8.069e-01
SD.log_k_cyan_bound_free 0.28537 6.809e-02 5.027e-01
SD.log_k_JCZ38 0.74846 2.305e-01 1.266e+00
SD.log_k_J9Z38 0.86077 2.713e-01 1.450e+00
SD.log_k_JSE76 0.97613 3.030e-01 1.649e+00
SD.f_cyan_ilr_1 0.38994 1.382e-01 6.417e-01
SD.f_cyan_ilr_2 0.82869 3.917e-02 1.618e+00
SD.f_JCZ38_qlogis 1.05000 -2.808e-02 2.128e+00
SD.f_JSE76_qlogis 0.44681 -3.985e+05 3.985e+05
Correlation:
cyn_f_0 lg_k_c_ lg_k_cyn_f_ lg_k_cyn_b_ l__JCZ3 l__J9Z3
log_k_cyan_free 0.0936
log_k_cyan_free_bound 0.1302 0.1627
log_k_cyan_bound_free 0.0029 0.0525 0.5181
log_k_JCZ38 -0.0116 -0.0077 -0.0430 -0.0236
log_k_J9Z38 -0.0192 -0.0077 -0.0048 0.0229 -0.0005
log_k_JSE76 0.0007 -0.0020 -0.0134 -0.0072 0.1225 -0.0016
f_cyan_ilr_1 -0.0118 -0.0027 -0.0132 -0.0118 0.0127 -0.0505
f_cyan_ilr_2 -0.4643 -0.0762 -0.1245 0.0137 0.0497 0.1003
f_JCZ38_qlogis 0.0710 0.0371 0.1826 0.0925 -0.0869 -0.0130
f_JSE76_qlogis -0.0367 -0.0270 -0.2274 -0.1865 0.1244 0.0098
l__JSE7 f_cy__1 f_cy__2 f_JCZ38
log_k_cyan_free
log_k_cyan_free_bound
log_k_cyan_bound_free
log_k_JCZ38
log_k_J9Z38
log_k_JSE76
f_cyan_ilr_1 0.0036
f_cyan_ilr_2 0.0050 -0.0201
f_JCZ38_qlogis 0.0142 -0.0529 -0.2698
f_JSE76_qlogis 0.0064 0.0345 0.2015 -0.7058
Random effects:
est. lower upper
SD.log_k_cyan_free 0.4967 1.865e-01 8.069e-01
SD.log_k_cyan_bound_free 0.2854 6.809e-02 5.027e-01
SD.log_k_JCZ38 0.7485 2.305e-01 1.266e+00
SD.log_k_J9Z38 0.8608 2.713e-01 1.450e+00
SD.log_k_JSE76 0.9761 3.030e-01 1.649e+00
SD.f_cyan_ilr_1 0.3899 1.382e-01 6.417e-01
SD.f_cyan_ilr_2 0.8287 3.917e-02 1.618e+00
SD.f_JCZ38_qlogis 1.0500 -2.808e-02 2.128e+00
SD.f_JSE76_qlogis 0.4468 -3.985e+05 3.985e+05
Variance model:
est. lower upper
a.1 2.07082 1.87680 2.26483
b.1 0.06227 0.05098 0.07355
Backtransformed parameters:
est. lower upper
cyan_free_0 1.007e+02 9.873e+01 102.72898
k_cyan_free 4.091e-02 2.623e-02 0.06382
k_cyan_free_bound 3.221e-02 2.537e-02 0.04090
k_cyan_bound_free 2.165e-02 1.557e-02 0.03011
k_JCZ38 8.122e-02 3.975e-02 0.16594
k_J9Z38 6.569e-03 2.954e-03 0.01461
k_JSE76 3.886e-02 1.556e-02 0.09703
f_cyan_free_to_JCZ38 6.785e-01 6.102e-01 0.97309
f_cyan_free_to_J9Z38 2.498e-01 6.102e-01 0.97309
f_JCZ38_to_JSE76 9.452e-01 7.392e-01 0.99056
f_JSE76_to_JCZ38 9.998e-01 5.580e-279 1.00000
Estimated Eigenvalues of SFORB model(s):
cyan_b1 cyan_b2 cyan_g
0.08426 0.01051 0.41220
Resulting formation fractions:
ff
cyan_free_JCZ38 0.6784541
cyan_free_J9Z38 0.2498405
cyan_free_sink 0.0717054
cyan_free 1.0000000
JCZ38_JSE76 0.9452043
JCZ38_sink 0.0547957
JSE76_JCZ38 0.9998226
JSE76_sink 0.0001774
Estimated disappearance times:
DT50 DT90 DT50back DT50_cyan_b1 DT50_cyan_b2
cyan 25.237 168.51 50.73 8.226 65.95
JCZ38 8.535 28.35 NA NA NA
J9Z38 105.517 350.52 NA NA NA
JSE76 17.837 59.25 NA NA NA
Pathway 2, refined fits
saemix version used for fitting: 3.3
mkin version used for pre-fitting: 1.2.10
R version used for fitting: 4.5.0
Date of fit: Mon May 12 21:04:10 2025
Date of summary: Mon May 12 21:06:22 2025
Equations:
d_cyan/dt = - (alpha/beta) * 1/((time/beta) + 1) * cyan
d_JCZ38/dt = + f_cyan_to_JCZ38 * (alpha/beta) * 1/((time/beta) + 1) *
cyan - k_JCZ38 * JCZ38 + f_JSE76_to_JCZ38 * k_JSE76 * JSE76
d_J9Z38/dt = + f_cyan_to_J9Z38 * (alpha/beta) * 1/((time/beta) + 1) *
cyan - k_J9Z38 * J9Z38
d_JSE76/dt = + f_JCZ38_to_JSE76 * k_JCZ38 * JCZ38 - k_JSE76 * JSE76
Data:
433 observations of 4 variable(s) grouped in 5 datasets
Model predictions using solution type deSolve
Fitted in 786.038 s
Using 300, 100 iterations and 10 chains
Variance model: Two-component variance function
Starting values for degradation parameters:
cyan_0 log_k_JCZ38 log_k_J9Z38 log_k_JSE76 f_cyan_ilr_1
102.4477 -1.8631 -5.1087 -2.5114 0.6826
f_cyan_ilr_2 f_JCZ38_qlogis f_JSE76_qlogis log_alpha log_beta
4.7944 15.9616 13.1566 -0.1564 2.9781
Fixed degradation parameter values:
None
Starting values for random effects (square root of initial entries in omega):
cyan_0 log_k_JCZ38 log_k_J9Z38 log_k_JSE76 f_cyan_ilr_1
cyan_0 7.701 0.000 0.000 0.000 0.0000
log_k_JCZ38 0.000 1.448 0.000 0.000 0.0000
log_k_J9Z38 0.000 0.000 1.724 0.000 0.0000
log_k_JSE76 0.000 0.000 0.000 3.659 0.0000
f_cyan_ilr_1 0.000 0.000 0.000 0.000 0.6356
f_cyan_ilr_2 0.000 0.000 0.000 0.000 0.0000
f_JCZ38_qlogis 0.000 0.000 0.000 0.000 0.0000
f_JSE76_qlogis 0.000 0.000 0.000 0.000 0.0000
log_alpha 0.000 0.000 0.000 0.000 0.0000
log_beta 0.000 0.000 0.000 0.000 0.0000
f_cyan_ilr_2 f_JCZ38_qlogis f_JSE76_qlogis log_alpha log_beta
cyan_0 0.00 0.00 0.00 0.0000 0.0000
log_k_JCZ38 0.00 0.00 0.00 0.0000 0.0000
log_k_J9Z38 0.00 0.00 0.00 0.0000 0.0000
log_k_JSE76 0.00 0.00 0.00 0.0000 0.0000
f_cyan_ilr_1 0.00 0.00 0.00 0.0000 0.0000
f_cyan_ilr_2 10.32 0.00 0.00 0.0000 0.0000
f_JCZ38_qlogis 0.00 12.23 0.00 0.0000 0.0000
f_JSE76_qlogis 0.00 0.00 14.99 0.0000 0.0000
log_alpha 0.00 0.00 0.00 0.3924 0.0000
log_beta 0.00 0.00 0.00 0.0000 0.5639
Starting values for error model parameters:
a.1 b.1
1 1
Results:
Likelihood computed by importance sampling
AIC BIC logLik
2249 2242 -1106
Optimised parameters:
est. lower upper
cyan_0 101.24524 NA NA
log_k_JCZ38 -2.85375 NA NA
log_k_J9Z38 -5.07729 NA NA
log_k_JSE76 -3.53511 NA NA
f_cyan_ilr_1 0.67478 NA NA
f_cyan_ilr_2 0.97152 NA NA
f_JCZ38_qlogis 213.48001 NA NA
f_JSE76_qlogis 2.02040 NA NA
log_alpha -0.11041 NA NA
log_beta 3.06575 NA NA
a.1 2.05279 1.85495 2.2506
b.1 0.07116 0.05912 0.0832
SD.log_k_JCZ38 1.21713 0.44160 1.9927
SD.log_k_J9Z38 0.88268 0.27541 1.4900
SD.log_k_JSE76 0.59452 0.15005 1.0390
SD.f_cyan_ilr_1 0.35370 0.12409 0.5833
SD.f_cyan_ilr_2 0.78186 0.18547 1.3782
SD.log_alpha 0.27781 0.08168 0.4739
SD.log_beta 0.32608 0.06490 0.5873
Correlation is not available
Random effects:
est. lower upper
SD.log_k_JCZ38 1.2171 0.44160 1.9927
SD.log_k_J9Z38 0.8827 0.27541 1.4900
SD.log_k_JSE76 0.5945 0.15005 1.0390
SD.f_cyan_ilr_1 0.3537 0.12409 0.5833
SD.f_cyan_ilr_2 0.7819 0.18547 1.3782
SD.log_alpha 0.2778 0.08168 0.4739
SD.log_beta 0.3261 0.06490 0.5873
Variance model:
est. lower upper
a.1 2.05279 1.85495 2.2506
b.1 0.07116 0.05912 0.0832
Backtransformed parameters:
est. lower upper
cyan_0 1.012e+02 NA NA
k_JCZ38 5.763e-02 NA NA
k_J9Z38 6.237e-03 NA NA
k_JSE76 2.916e-02 NA NA
f_cyan_to_JCZ38 6.354e-01 NA NA
f_cyan_to_J9Z38 2.447e-01 NA NA
f_JCZ38_to_JSE76 1.000e+00 NA NA
f_JSE76_to_JCZ38 8.829e-01 NA NA
alpha 8.955e-01 NA NA
beta 2.145e+01 NA NA
Resulting formation fractions:
ff
cyan_JCZ38 0.6354
cyan_J9Z38 0.2447
cyan_sink 0.1200
JCZ38_JSE76 1.0000
JCZ38_sink 0.0000
JSE76_JCZ38 0.8829
JSE76_sink 0.1171
Estimated disappearance times:
DT50 DT90 DT50back
cyan 25.07 259.21 78.03
JCZ38 12.03 39.96 NA
J9Z38 111.14 369.19 NA
JSE76 23.77 78.98 NA
saemix version used for fitting: 3.3
mkin version used for pre-fitting: 1.2.10
R version used for fitting: 4.5.0
Date of fit: Mon May 12 21:05:56 2025
Date of summary: Mon May 12 21:06:22 2025
Equations:
d_cyan/dt = - ((k1 * g * exp(-k1 * time) + k2 * (1 - g) * exp(-k2 *
time)) / (g * exp(-k1 * time) + (1 - g) * exp(-k2 * time)))
* cyan
d_JCZ38/dt = + f_cyan_to_JCZ38 * ((k1 * g * exp(-k1 * time) + k2 * (1 -
g) * exp(-k2 * time)) / (g * exp(-k1 * time) + (1 - g) *
exp(-k2 * time))) * cyan - k_JCZ38 * JCZ38 +
f_JSE76_to_JCZ38 * k_JSE76 * JSE76
d_J9Z38/dt = + f_cyan_to_J9Z38 * ((k1 * g * exp(-k1 * time) + k2 * (1 -
g) * exp(-k2 * time)) / (g * exp(-k1 * time) + (1 - g) *
exp(-k2 * time))) * cyan - k_J9Z38 * J9Z38
d_JSE76/dt = + f_JCZ38_to_JSE76 * k_JCZ38 * JCZ38 - k_JSE76 * JSE76
Data:
433 observations of 4 variable(s) grouped in 5 datasets
Model predictions using solution type deSolve
Fitted in 892.139 s
Using 300, 100 iterations and 10 chains
Variance model: Constant variance
Starting values for degradation parameters:
cyan_0 log_k_JCZ38 log_k_J9Z38 log_k_JSE76 f_cyan_ilr_1
102.4380 -2.3107 -5.3123 -3.7120 0.6757
f_cyan_ilr_2 f_JCZ38_qlogis f_JSE76_qlogis log_k1 log_k2
1.1439 13.1194 12.3492 -1.9317 -4.4557
g_qlogis
-0.5644
Fixed degradation parameter values:
None
Starting values for random effects (square root of initial entries in omega):
cyan_0 log_k_JCZ38 log_k_J9Z38 log_k_JSE76 f_cyan_ilr_1
cyan_0 4.591 0.0000 0.000 0.0 0.0000
log_k_JCZ38 0.000 0.7966 0.000 0.0 0.0000
log_k_J9Z38 0.000 0.0000 1.561 0.0 0.0000
log_k_JSE76 0.000 0.0000 0.000 0.8 0.0000
f_cyan_ilr_1 0.000 0.0000 0.000 0.0 0.6349
f_cyan_ilr_2 0.000 0.0000 0.000 0.0 0.0000
f_JCZ38_qlogis 0.000 0.0000 0.000 0.0 0.0000
f_JSE76_qlogis 0.000 0.0000 0.000 0.0 0.0000
log_k1 0.000 0.0000 0.000 0.0 0.0000
log_k2 0.000 0.0000 0.000 0.0 0.0000
g_qlogis 0.000 0.0000 0.000 0.0 0.0000
f_cyan_ilr_2 f_JCZ38_qlogis f_JSE76_qlogis log_k1 log_k2
cyan_0 0.000 0.00 0.00 0.000 0.0000
log_k_JCZ38 0.000 0.00 0.00 0.000 0.0000
log_k_J9Z38 0.000 0.00 0.00 0.000 0.0000
log_k_JSE76 0.000 0.00 0.00 0.000 0.0000
f_cyan_ilr_1 0.000 0.00 0.00 0.000 0.0000
f_cyan_ilr_2 1.797 0.00 0.00 0.000 0.0000
f_JCZ38_qlogis 0.000 13.86 0.00 0.000 0.0000
f_JSE76_qlogis 0.000 0.00 13.91 0.000 0.0000
log_k1 0.000 0.00 0.00 1.106 0.0000
log_k2 0.000 0.00 0.00 0.000 0.6141
g_qlogis 0.000 0.00 0.00 0.000 0.0000
g_qlogis
cyan_0 0.000
log_k_JCZ38 0.000
log_k_J9Z38 0.000
log_k_JSE76 0.000
f_cyan_ilr_1 0.000
f_cyan_ilr_2 0.000
f_JCZ38_qlogis 0.000
f_JSE76_qlogis 0.000
log_k1 0.000
log_k2 0.000
g_qlogis 1.595
Starting values for error model parameters:
a.1
1
Results:
Likelihood computed by importance sampling
AIC BIC logLik
2282 2274 -1121
Optimised parameters:
est. lower upper
cyan_0 102.6036 NA NA
log_k_JCZ38 -2.9348 NA NA
log_k_J9Z38 -5.1617 NA NA
log_k_JSE76 -3.6396 NA NA
f_cyan_ilr_1 0.6991 NA NA
f_cyan_ilr_2 0.6341 NA NA
f_JCZ38_qlogis 4232.3011 NA NA
f_JSE76_qlogis 1.9658 NA NA
log_k1 -1.9503 NA NA
log_k2 -4.4745 NA NA
g_qlogis -0.4967 NA NA
a.1 2.7461 2.59274 2.8994
SD.log_k_JCZ38 1.3178 0.47602 2.1596
SD.log_k_J9Z38 0.7022 0.15061 1.2538
SD.log_k_JSE76 0.6566 0.15613 1.1570
SD.f_cyan_ilr_1 0.3409 0.11666 0.5652
SD.f_cyan_ilr_2 0.4385 0.09482 0.7821
SD.log_k1 0.7381 0.25599 1.2202
SD.log_k2 0.5133 0.18152 0.8450
SD.g_qlogis 0.9866 0.35681 1.6164
Correlation is not available
Random effects:
est. lower upper
SD.log_k_JCZ38 1.3178 0.47602 2.1596
SD.log_k_J9Z38 0.7022 0.15061 1.2538
SD.log_k_JSE76 0.6566 0.15613 1.1570
SD.f_cyan_ilr_1 0.3409 0.11666 0.5652
SD.f_cyan_ilr_2 0.4385 0.09482 0.7821
SD.log_k1 0.7381 0.25599 1.2202
SD.log_k2 0.5133 0.18152 0.8450
SD.g_qlogis 0.9866 0.35681 1.6164
Variance model:
est. lower upper
a.1 2.746 2.593 2.899
Backtransformed parameters:
est. lower upper
cyan_0 1.026e+02 NA NA
k_JCZ38 5.314e-02 NA NA
k_J9Z38 5.732e-03 NA NA
k_JSE76 2.626e-02 NA NA
f_cyan_to_JCZ38 6.051e-01 NA NA
f_cyan_to_J9Z38 2.251e-01 NA NA
f_JCZ38_to_JSE76 1.000e+00 NA NA
f_JSE76_to_JCZ38 8.772e-01 NA NA
k1 1.422e-01 NA NA
k2 1.140e-02 NA NA
g 3.783e-01 NA NA
Resulting formation fractions:
ff
cyan_JCZ38 0.6051
cyan_J9Z38 0.2251
cyan_sink 0.1698
JCZ38_JSE76 1.0000
JCZ38_sink 0.0000
JSE76_JCZ38 0.8772
JSE76_sink 0.1228
Estimated disappearance times:
DT50 DT90 DT50back DT50_k1 DT50_k2
cyan 22.05 160.35 48.27 4.873 60.83
JCZ38 13.04 43.33 NA NA NA
J9Z38 120.93 401.73 NA NA NA
JSE76 26.39 87.68 NA NA NA
saemix version used for fitting: 3.3
mkin version used for pre-fitting: 1.2.10
R version used for fitting: 4.5.0
Date of fit: Mon May 12 21:06:02 2025
Date of summary: Mon May 12 21:06:22 2025
Equations:
d_cyan/dt = - ((k1 * g * exp(-k1 * time) + k2 * (1 - g) * exp(-k2 *
time)) / (g * exp(-k1 * time) + (1 - g) * exp(-k2 * time)))
* cyan
d_JCZ38/dt = + f_cyan_to_JCZ38 * ((k1 * g * exp(-k1 * time) + k2 * (1 -
g) * exp(-k2 * time)) / (g * exp(-k1 * time) + (1 - g) *
exp(-k2 * time))) * cyan - k_JCZ38 * JCZ38 +
f_JSE76_to_JCZ38 * k_JSE76 * JSE76
d_J9Z38/dt = + f_cyan_to_J9Z38 * ((k1 * g * exp(-k1 * time) + k2 * (1 -
g) * exp(-k2 * time)) / (g * exp(-k1 * time) + (1 - g) *
exp(-k2 * time))) * cyan - k_J9Z38 * J9Z38
d_JSE76/dt = + f_JCZ38_to_JSE76 * k_JCZ38 * JCZ38 - k_JSE76 * JSE76
Data:
433 observations of 4 variable(s) grouped in 5 datasets
Model predictions using solution type deSolve
Fitted in 898.534 s
Using 300, 100 iterations and 10 chains
Variance model: Two-component variance function
Starting values for degradation parameters:
cyan_0 log_k_JCZ38 log_k_J9Z38 log_k_JSE76 f_cyan_ilr_1
101.7393 -1.4493 -5.0118 -2.1269 0.6720
f_cyan_ilr_2 f_JCZ38_qlogis f_JSE76_qlogis log_k1 log_k2
7.3362 13.4423 13.2659 -2.0061 -4.5527
g_qlogis
-0.5806
Fixed degradation parameter values:
None
Starting values for random effects (square root of initial entries in omega):
cyan_0 log_k_JCZ38 log_k_J9Z38 log_k_JSE76 f_cyan_ilr_1
cyan_0 5.604 0.00 0.000 0.000 0.0000
log_k_JCZ38 0.000 2.77 0.000 0.000 0.0000
log_k_J9Z38 0.000 0.00 1.662 0.000 0.0000
log_k_JSE76 0.000 0.00 0.000 5.021 0.0000
f_cyan_ilr_1 0.000 0.00 0.000 0.000 0.6519
f_cyan_ilr_2 0.000 0.00 0.000 0.000 0.0000
f_JCZ38_qlogis 0.000 0.00 0.000 0.000 0.0000
f_JSE76_qlogis 0.000 0.00 0.000 0.000 0.0000
log_k1 0.000 0.00 0.000 0.000 0.0000
log_k2 0.000 0.00 0.000 0.000 0.0000
g_qlogis 0.000 0.00 0.000 0.000 0.0000
f_cyan_ilr_2 f_JCZ38_qlogis f_JSE76_qlogis log_k1 log_k2
cyan_0 0.00 0.00 0.00 0.0000 0.0000
log_k_JCZ38 0.00 0.00 0.00 0.0000 0.0000
log_k_J9Z38 0.00 0.00 0.00 0.0000 0.0000
log_k_JSE76 0.00 0.00 0.00 0.0000 0.0000
f_cyan_ilr_1 0.00 0.00 0.00 0.0000 0.0000
f_cyan_ilr_2 13.37 0.00 0.00 0.0000 0.0000
f_JCZ38_qlogis 0.00 14.21 0.00 0.0000 0.0000
f_JSE76_qlogis 0.00 0.00 14.58 0.0000 0.0000
log_k1 0.00 0.00 0.00 0.8453 0.0000
log_k2 0.00 0.00 0.00 0.0000 0.5969
g_qlogis 0.00 0.00 0.00 0.0000 0.0000
g_qlogis
cyan_0 0.00
log_k_JCZ38 0.00
log_k_J9Z38 0.00
log_k_JSE76 0.00
f_cyan_ilr_1 0.00
f_cyan_ilr_2 0.00
f_JCZ38_qlogis 0.00
f_JSE76_qlogis 0.00
log_k1 0.00
log_k2 0.00
g_qlogis 1.69
Starting values for error model parameters:
a.1 b.1
1 1
Results:
Likelihood computed by importance sampling
AIC BIC logLik
2237 2229 -1099
Optimised parameters:
est. lower upper
cyan_0 101.00243 NA NA
log_k_JCZ38 -2.80828 NA NA
log_k_J9Z38 -5.04449 NA NA
log_k_JSE76 -3.66981 NA NA
f_cyan_ilr_1 0.72564 NA NA
f_cyan_ilr_2 1.37978 NA NA
f_JCZ38_qlogis 1.98726 NA NA
f_JSE76_qlogis 414.80884 NA NA
log_k1 -2.38601 NA NA
log_k2 -4.63632 NA NA
g_qlogis -0.33920 NA NA
a.1 2.10837 1.91261 2.30413
b.1 0.06223 0.05085 0.07361
SD.log_k_JCZ38 1.30902 0.48128 2.13675
SD.log_k_J9Z38 0.83882 0.25790 1.41974
SD.log_k_JSE76 0.58104 0.14201 1.02008
SD.f_cyan_ilr_1 0.35421 0.12398 0.58443
SD.f_cyan_ilr_2 0.79373 0.12007 1.46739
SD.log_k2 0.27476 0.08557 0.46394
SD.g_qlogis 0.96170 0.35463 1.56878
Correlation is not available
Random effects:
est. lower upper
SD.log_k_JCZ38 1.3090 0.48128 2.1367
SD.log_k_J9Z38 0.8388 0.25790 1.4197
SD.log_k_JSE76 0.5810 0.14201 1.0201
SD.f_cyan_ilr_1 0.3542 0.12398 0.5844
SD.f_cyan_ilr_2 0.7937 0.12007 1.4674
SD.log_k2 0.2748 0.08557 0.4639
SD.g_qlogis 0.9617 0.35463 1.5688
Variance model:
est. lower upper
a.1 2.10837 1.91261 2.30413
b.1 0.06223 0.05085 0.07361
Backtransformed parameters:
est. lower upper
cyan_0 1.010e+02 NA NA
k_JCZ38 6.031e-02 NA NA
k_J9Z38 6.445e-03 NA NA
k_JSE76 2.548e-02 NA NA
f_cyan_to_JCZ38 6.808e-01 NA NA
f_cyan_to_J9Z38 2.440e-01 NA NA
f_JCZ38_to_JSE76 8.795e-01 NA NA
f_JSE76_to_JCZ38 1.000e+00 NA NA
k1 9.200e-02 NA NA
k2 9.693e-03 NA NA
g 4.160e-01 NA NA
Resulting formation fractions:
ff
cyan_JCZ38 0.68081
cyan_J9Z38 0.24398
cyan_sink 0.07521
JCZ38_JSE76 0.87945
JCZ38_sink 0.12055
JSE76_JCZ38 1.00000
JSE76_sink 0.00000
Estimated disappearance times:
DT50 DT90 DT50back DT50_k1 DT50_k2
cyan 25.00 182.05 54.8 7.535 71.51
JCZ38 11.49 38.18 NA NA NA
J9Z38 107.55 357.28 NA NA NA
JSE76 27.20 90.36 NA NA NA
saemix version used for fitting: 3.3
mkin version used for pre-fitting: 1.2.10
R version used for fitting: 4.5.0
Date of fit: Mon May 12 21:05:52 2025
Date of summary: Mon May 12 21:06:22 2025
Equations:
d_cyan_free/dt = - k_cyan_free * cyan_free - k_cyan_free_bound *
cyan_free + k_cyan_bound_free * cyan_bound
d_cyan_bound/dt = + k_cyan_free_bound * cyan_free - k_cyan_bound_free *
cyan_bound
d_JCZ38/dt = + f_cyan_free_to_JCZ38 * k_cyan_free * cyan_free - k_JCZ38
* JCZ38 + f_JSE76_to_JCZ38 * k_JSE76 * JSE76
d_J9Z38/dt = + f_cyan_free_to_J9Z38 * k_cyan_free * cyan_free - k_J9Z38
* J9Z38
d_JSE76/dt = + f_JCZ38_to_JSE76 * k_JCZ38 * JCZ38 - k_JSE76 * JSE76
Data:
433 observations of 4 variable(s) grouped in 5 datasets
Model predictions using solution type deSolve
Fitted in 888.333 s
Using 300, 100 iterations and 10 chains
Variance model: Constant variance
Starting values for degradation parameters:
cyan_free_0 log_k_cyan_free log_k_cyan_free_bound
102.4395 -2.7673 -2.8942
log_k_cyan_bound_free log_k_JCZ38 log_k_J9Z38
-3.6201 -2.3107 -5.3123
log_k_JSE76 f_cyan_ilr_1 f_cyan_ilr_2
-3.7120 0.6754 1.1448
f_JCZ38_qlogis f_JSE76_qlogis
14.8408 15.4734
Fixed degradation parameter values:
None
Starting values for random effects (square root of initial entries in omega):
cyan_free_0 log_k_cyan_free log_k_cyan_free_bound
cyan_free_0 4.589 0.0000 0.00
log_k_cyan_free 0.000 0.4849 0.00
log_k_cyan_free_bound 0.000 0.0000 1.62
log_k_cyan_bound_free 0.000 0.0000 0.00
log_k_JCZ38 0.000 0.0000 0.00
log_k_J9Z38 0.000 0.0000 0.00
log_k_JSE76 0.000 0.0000 0.00
f_cyan_ilr_1 0.000 0.0000 0.00
f_cyan_ilr_2 0.000 0.0000 0.00
f_JCZ38_qlogis 0.000 0.0000 0.00
f_JSE76_qlogis 0.000 0.0000 0.00
log_k_cyan_bound_free log_k_JCZ38 log_k_J9Z38 log_k_JSE76
cyan_free_0 0.000 0.0000 0.000 0.0
log_k_cyan_free 0.000 0.0000 0.000 0.0
log_k_cyan_free_bound 0.000 0.0000 0.000 0.0
log_k_cyan_bound_free 1.197 0.0000 0.000 0.0
log_k_JCZ38 0.000 0.7966 0.000 0.0
log_k_J9Z38 0.000 0.0000 1.561 0.0
log_k_JSE76 0.000 0.0000 0.000 0.8
f_cyan_ilr_1 0.000 0.0000 0.000 0.0
f_cyan_ilr_2 0.000 0.0000 0.000 0.0
f_JCZ38_qlogis 0.000 0.0000 0.000 0.0
f_JSE76_qlogis 0.000 0.0000 0.000 0.0
f_cyan_ilr_1 f_cyan_ilr_2 f_JCZ38_qlogis f_JSE76_qlogis
cyan_free_0 0.0000 0.000 0.0 0.00
log_k_cyan_free 0.0000 0.000 0.0 0.00
log_k_cyan_free_bound 0.0000 0.000 0.0 0.00
log_k_cyan_bound_free 0.0000 0.000 0.0 0.00
log_k_JCZ38 0.0000 0.000 0.0 0.00
log_k_J9Z38 0.0000 0.000 0.0 0.00
log_k_JSE76 0.0000 0.000 0.0 0.00
f_cyan_ilr_1 0.6349 0.000 0.0 0.00
f_cyan_ilr_2 0.0000 1.797 0.0 0.00
f_JCZ38_qlogis 0.0000 0.000 15.6 0.00
f_JSE76_qlogis 0.0000 0.000 0.0 17.52
Starting values for error model parameters:
a.1
1
Results:
Likelihood computed by importance sampling
AIC BIC logLik
2280 2272 -1120
Optimised parameters:
est. lower upper
cyan_free_0 102.6532 NA NA
log_k_cyan_free -2.8547 NA NA
log_k_cyan_free_bound -2.7004 NA NA
log_k_cyan_bound_free -3.5078 NA NA
log_k_JCZ38 -2.9255 NA NA
log_k_J9Z38 -5.1089 NA NA
log_k_JSE76 -3.6263 NA NA
f_cyan_ilr_1 0.6873 NA NA
f_cyan_ilr_2 0.6498 NA NA
f_JCZ38_qlogis 3624.2149 NA NA
f_JSE76_qlogis 1.9991 NA NA
a.1 2.7472 2.55559 2.9388
SD.log_k_cyan_free 0.3227 0.10296 0.5423
SD.log_k_cyan_free_bound 0.8757 0.29525 1.4562
SD.log_k_cyan_bound_free 0.6128 0.20220 1.0233
SD.log_k_JCZ38 1.3431 0.48474 2.2014
SD.log_k_J9Z38 0.6881 0.14714 1.2291
SD.log_k_JSE76 0.6461 0.15321 1.1390
SD.f_cyan_ilr_1 0.3361 0.11376 0.5585
SD.f_cyan_ilr_2 0.4286 0.08419 0.7730
Correlation is not available
Random effects:
est. lower upper
SD.log_k_cyan_free 0.3227 0.10296 0.5423
SD.log_k_cyan_free_bound 0.8757 0.29525 1.4562
SD.log_k_cyan_bound_free 0.6128 0.20220 1.0233
SD.log_k_JCZ38 1.3431 0.48474 2.2014
SD.log_k_J9Z38 0.6881 0.14714 1.2291
SD.log_k_JSE76 0.6461 0.15321 1.1390
SD.f_cyan_ilr_1 0.3361 0.11376 0.5585
SD.f_cyan_ilr_2 0.4286 0.08419 0.7730
Variance model:
est. lower upper
a.1 2.747 2.556 2.939
Backtransformed parameters:
est. lower upper
cyan_free_0 1.027e+02 NA NA
k_cyan_free 5.758e-02 NA NA
k_cyan_free_bound 6.718e-02 NA NA
k_cyan_bound_free 2.996e-02 NA NA
k_JCZ38 5.364e-02 NA NA
k_J9Z38 6.042e-03 NA NA
k_JSE76 2.662e-02 NA NA
f_cyan_free_to_JCZ38 6.039e-01 NA NA
f_cyan_free_to_J9Z38 2.285e-01 NA NA
f_JCZ38_to_JSE76 1.000e+00 NA NA
f_JSE76_to_JCZ38 8.807e-01 NA NA
Estimated Eigenvalues of SFORB model(s):
cyan_b1 cyan_b2 cyan_g
0.1426 0.0121 0.3484
Resulting formation fractions:
ff
cyan_free_JCZ38 0.6039
cyan_free_J9Z38 0.2285
cyan_free_sink 0.1676
cyan_free 1.0000
JCZ38_JSE76 1.0000
JCZ38_sink 0.0000
JSE76_JCZ38 0.8807
JSE76_sink 0.1193
Estimated disappearance times:
DT50 DT90 DT50back DT50_cyan_b1 DT50_cyan_b2
cyan 23.84 154.95 46.65 4.86 57.31
JCZ38 12.92 42.93 NA NA NA
J9Z38 114.71 381.07 NA NA NA
JSE76 26.04 86.51 NA NA NA
saemix version used for fitting: 3.3
mkin version used for pre-fitting: 1.2.10
R version used for fitting: 4.5.0
Date of fit: Mon May 12 21:06:21 2025
Date of summary: Mon May 12 21:06:22 2025
Equations:
d_cyan_free/dt = - k_cyan_free * cyan_free - k_cyan_free_bound *
cyan_free + k_cyan_bound_free * cyan_bound
d_cyan_bound/dt = + k_cyan_free_bound * cyan_free - k_cyan_bound_free *
cyan_bound
d_JCZ38/dt = + f_cyan_free_to_JCZ38 * k_cyan_free * cyan_free - k_JCZ38
* JCZ38 + f_JSE76_to_JCZ38 * k_JSE76 * JSE76
d_J9Z38/dt = + f_cyan_free_to_J9Z38 * k_cyan_free * cyan_free - k_J9Z38
* J9Z38
d_JSE76/dt = + f_JCZ38_to_JSE76 * k_JCZ38 * JCZ38 - k_JSE76 * JSE76
Data:
433 observations of 4 variable(s) grouped in 5 datasets
Model predictions using solution type deSolve
Fitted in 916.619 s
Using 300, 100 iterations and 10 chains
Variance model: Two-component variance function
Starting values for degradation parameters:
cyan_free_0 log_k_cyan_free log_k_cyan_free_bound
101.7511 -2.8370 -3.0162
log_k_cyan_bound_free log_k_JCZ38 log_k_J9Z38
-3.6600 -2.2988 -5.3129
log_k_JSE76 f_cyan_ilr_1 f_cyan_ilr_2
-3.6991 0.6722 4.8596
f_JCZ38_qlogis f_JSE76_qlogis
13.4678 14.2149
Fixed degradation parameter values:
None
Starting values for random effects (square root of initial entries in omega):
cyan_free_0 log_k_cyan_free log_k_cyan_free_bound
cyan_free_0 5.629 0.000 0.000
log_k_cyan_free 0.000 0.446 0.000
log_k_cyan_free_bound 0.000 0.000 1.449
log_k_cyan_bound_free 0.000 0.000 0.000
log_k_JCZ38 0.000 0.000 0.000
log_k_J9Z38 0.000 0.000 0.000
log_k_JSE76 0.000 0.000 0.000
f_cyan_ilr_1 0.000 0.000 0.000
f_cyan_ilr_2 0.000 0.000 0.000
f_JCZ38_qlogis 0.000 0.000 0.000
f_JSE76_qlogis 0.000 0.000 0.000
log_k_cyan_bound_free log_k_JCZ38 log_k_J9Z38 log_k_JSE76
cyan_free_0 0.000 0.0000 0.000 0.0000
log_k_cyan_free 0.000 0.0000 0.000 0.0000
log_k_cyan_free_bound 0.000 0.0000 0.000 0.0000
log_k_cyan_bound_free 1.213 0.0000 0.000 0.0000
log_k_JCZ38 0.000 0.7801 0.000 0.0000
log_k_J9Z38 0.000 0.0000 1.575 0.0000
log_k_JSE76 0.000 0.0000 0.000 0.8078
f_cyan_ilr_1 0.000 0.0000 0.000 0.0000
f_cyan_ilr_2 0.000 0.0000 0.000 0.0000
f_JCZ38_qlogis 0.000 0.0000 0.000 0.0000
f_JSE76_qlogis 0.000 0.0000 0.000 0.0000
f_cyan_ilr_1 f_cyan_ilr_2 f_JCZ38_qlogis f_JSE76_qlogis
cyan_free_0 0.0000 0.000 0.00 0.00
log_k_cyan_free 0.0000 0.000 0.00 0.00
log_k_cyan_free_bound 0.0000 0.000 0.00 0.00
log_k_cyan_bound_free 0.0000 0.000 0.00 0.00
log_k_JCZ38 0.0000 0.000 0.00 0.00
log_k_J9Z38 0.0000 0.000 0.00 0.00
log_k_JSE76 0.0000 0.000 0.00 0.00
f_cyan_ilr_1 0.6518 0.000 0.00 0.00
f_cyan_ilr_2 0.0000 9.981 0.00 0.00
f_JCZ38_qlogis 0.0000 0.000 14.26 0.00
f_JSE76_qlogis 0.0000 0.000 0.00 16.17
Starting values for error model parameters:
a.1 b.1
1 1
Results:
Likelihood computed by importance sampling
AIC BIC logLik
2241 2233 -1101
Optimised parameters:
est. lower upper
cyan_free_0 100.95469 NA NA
log_k_cyan_free -3.18706 NA NA
log_k_cyan_free_bound -3.38455 NA NA
log_k_cyan_bound_free -3.75788 NA NA
log_k_JCZ38 -2.77024 NA NA
log_k_J9Z38 -5.03665 NA NA
log_k_JSE76 -3.60289 NA NA
f_cyan_ilr_1 0.72263 NA NA
f_cyan_ilr_2 1.45352 NA NA
f_JCZ38_qlogis 2.00778 NA NA
f_JSE76_qlogis 941.58570 NA NA
a.1 2.11130 1.91479 2.30780
b.1 0.06299 0.05152 0.07445
SD.log_k_cyan_free 0.50098 0.18805 0.81390
SD.log_k_cyan_bound_free 0.31671 0.08467 0.54875
SD.log_k_JCZ38 1.25865 0.45932 2.05798
SD.log_k_J9Z38 0.86833 0.27222 1.46444
SD.log_k_JSE76 0.59325 0.14711 1.03940
SD.f_cyan_ilr_1 0.35705 0.12521 0.58890
SD.f_cyan_ilr_2 0.88541 0.13797 1.63286
Correlation is not available
Random effects:
est. lower upper
SD.log_k_cyan_free 0.5010 0.18805 0.8139
SD.log_k_cyan_bound_free 0.3167 0.08467 0.5487
SD.log_k_JCZ38 1.2587 0.45932 2.0580
SD.log_k_J9Z38 0.8683 0.27222 1.4644
SD.log_k_JSE76 0.5933 0.14711 1.0394
SD.f_cyan_ilr_1 0.3571 0.12521 0.5889
SD.f_cyan_ilr_2 0.8854 0.13797 1.6329
Variance model:
est. lower upper
a.1 2.11130 1.91479 2.30780
b.1 0.06299 0.05152 0.07445
Backtransformed parameters:
est. lower upper
cyan_free_0 1.010e+02 NA NA
k_cyan_free 4.129e-02 NA NA
k_cyan_free_bound 3.389e-02 NA NA
k_cyan_bound_free 2.333e-02 NA NA
k_JCZ38 6.265e-02 NA NA
k_J9Z38 6.495e-03 NA NA
k_JSE76 2.724e-02 NA NA
f_cyan_free_to_JCZ38 6.844e-01 NA NA
f_cyan_free_to_J9Z38 2.463e-01 NA NA
f_JCZ38_to_JSE76 8.816e-01 NA NA
f_JSE76_to_JCZ38 1.000e+00 NA NA
Estimated Eigenvalues of SFORB model(s):
cyan_b1 cyan_b2 cyan_g
0.08751 0.01101 0.39586
Resulting formation fractions:
ff
cyan_free_JCZ38 0.68444
cyan_free_J9Z38 0.24633
cyan_free_sink 0.06923
cyan_free 1.00000
JCZ38_JSE76 0.88161
JCZ38_sink 0.11839
JSE76_JCZ38 1.00000
JSE76_sink 0.00000
Estimated disappearance times:
DT50 DT90 DT50back DT50_cyan_b1 DT50_cyan_b2
cyan 25.36 163.36 49.18 7.921 62.95
JCZ38 11.06 36.75 NA NA NA
J9Z38 106.71 354.49 NA NA NA
JSE76 25.44 84.51 NA NA NA
Session info
R version 4.5.0 (2025-04-11)
Platform: x86_64-pc-linux-gnu
Running under: Debian GNU/Linux 12 (bookworm)
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/blas/libblas.so.3.11.0
LAPACK: /usr/lib/x86_64-linux-gnu/lapack/liblapack.so.3.11.0 LAPACK version 3.11.0
locale:
[1] LC_CTYPE=de_DE.UTF-8 LC_NUMERIC=C
[3] LC_TIME=de_DE.UTF-8 LC_COLLATE=de_DE.UTF-8
[5] LC_MONETARY=de_DE.UTF-8 LC_MESSAGES=de_DE.UTF-8
[7] LC_PAPER=de_DE.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=de_DE.UTF-8 LC_IDENTIFICATION=C
time zone: Europe/Berlin
tzcode source: system (glibc)
attached base packages:
[1] parallel stats graphics grDevices utils datasets methods
[8] base
other attached packages:
[1] rmarkdown_2.29 nvimcom_0.9-167 saemix_3.3 npde_3.5
[5] knitr_1.49 mkin_1.2.10
loaded via a namespace (and not attached):
[1] sass_0.4.9 generics_0.1.3 lattice_0.22-6 digest_0.6.37
[5] magrittr_2.0.3 evaluate_1.0.3 grid_4.5.0 fastmap_1.2.0
[9] cellranger_1.1.0 jsonlite_1.9.0 processx_3.8.6 pkgbuild_1.4.6
[13] deSolve_1.40 mclust_6.1.1 ps_1.9.0 gridExtra_2.3
[17] scales_1.3.0 codetools_0.2-20 textshaping_1.0.0 jquerylib_0.1.4
[21] cli_3.6.4 rlang_1.1.5 munsell_0.5.1 cachem_1.1.0
[25] yaml_2.3.10 inline_0.3.21 tools_4.5.0 dplyr_1.1.4
[29] colorspace_2.1-1 ggplot2_3.5.1 vctrs_0.6.5 R6_2.6.1
[33] zoo_1.8-13 lifecycle_1.0.4 fs_1.6.5 htmlwidgets_1.6.4
[37] MASS_7.3-65 ragg_1.3.3 pkgconfig_2.0.3 desc_1.4.3
[41] callr_3.7.6 pkgdown_2.1.1 pillar_1.10.1 bslib_0.9.0
[45] gtable_0.3.6 glue_1.8.0 systemfonts_1.2.1 xfun_0.51
[49] tibble_3.2.1 lmtest_0.9-40 tidyselect_1.2.1 htmltools_0.5.8.1
[53] nlme_3.1-168 compiler_4.5.0 readxl_1.4.4